Introgression of a major QTL from an inferior into a superior population using genomic selection
- PMID: 19635140
- PMCID: PMC2731732
- DOI: 10.1186/1297-9686-41-38
Introgression of a major QTL from an inferior into a superior population using genomic selection
Abstract
Background: Selection schemes aiming at introgressing genetic material from a donor into a recipient line may be performed by backcross-breeding programs combined with selection to preserve the favourable characteristics of the donor population. This stochastic simulation study investigated whether genomic selection can be effective in preserving a major quantitative trait locus (QTL) allele from a donor line during the backcrossing phase.
Methods: In a simulation study, two fish populations were generated: a recipient line selected for a production trait and a donor line characterized by an enhanced level of disease resistance. Both traits were polygenic, but one major QTL affecting disease resistance was segregating only within the donor line. Backcrossing was combined with three types of selection (for total merit index) among the crossbred individuals: classical selection, genomic selection using genome-wide dense marker maps, and gene-assisted genomic selection. It was assumed that production could be observed directly on the selection candidates, while disease resistance had to be inferred from tested sibs of the selection candidates.
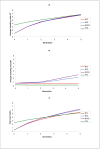
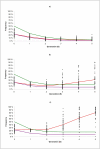
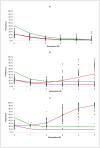
Results: Classical selection was inefficient in preserving the target QTL through the backcrossing phase. In contrast, genomic selection (without specific knowledge of the target QTL) was usually effective in preserving the target QTL, and had higher genetic response to selection, especially for disease resistance. Compared with pure genomic selection, gene-assisted selection had an advantage with respect to disease resistance (28-40% increase in genetic gain) and acted as an extra precaution against loss of the target QTL. However, for total merit index the advantage of gene-assisted genomic selection over genomic selection was lower (4-5% increase in genetic gain).
Conclusion: Substantial differences between introgression programs using classical and genomic selection were observed, and the former was generally inferior with respect to both genetic gain and the ability to preserve the target QTL. Combining genomic selection with gene-assisted selection for the target QTL acted as an extra precaution against loss of the target QTL and gave additional genetic gain for disease resistance. However, the effect on total merit index was limited.
Figures
Similar articles
-
Incorporating desirable genetic characteristics from an inferior into a superior population using genomic selection.Genetics. 2009 Feb;181(2):737-45. doi: 10.1534/genetics.108.098160. Epub 2008 Dec 1. Genetics. 2009. PMID: 19047412 Free PMC article.
-
Maximizing genetic gain over multiple generations with quantitative trait locus selection and control of inbreeding.J Anim Sci. 2004 May;82(5):1305-14. doi: 10.2527/2004.8251305x. J Anim Sci. 2004. PMID: 15144069
-
Comparison of different foreground and background selection methods in marker-assisted introgression.Yi Chuan Xue Bao. 2006 Dec;33(12):1073-80. doi: 10.1016/S0379-4172(06)60144-3. Yi Chuan Xue Bao. 2006. PMID: 17185166
-
Simultaneous selection of major and minor genes: use of QTL to increase selection efficiency of coleoptile length of wheat (Triticum aestivum L.).Theor Appl Genet. 2009 Jun;119(1):65-74. doi: 10.1007/s00122-009-1017-2. Epub 2009 Apr 10. Theor Appl Genet. 2009. PMID: 19360392
-
Genetics and genomics of disease resistance in salmonid species.Front Genet. 2014 Nov 26;5:415. doi: 10.3389/fgene.2014.00415. eCollection 2014. Front Genet. 2014. PMID: 25505486 Free PMC article. Review.
Cited by
-
Utilization of farm animal genetic resources in a changing agro-ecological environment in the Nordic countries.Front Genet. 2015 Feb 25;6:52. doi: 10.3389/fgene.2015.00052. eCollection 2015. Front Genet. 2015. PMID: 25767477 Free PMC article.
-
Benefit of Introgression Depends on Level of Genetic Trait Variation in Cereal Breeding Programmes.Front Plant Sci. 2022 Jun 15;13:786452. doi: 10.3389/fpls.2022.786452. eCollection 2022. Front Plant Sci. 2022. PMID: 35783964 Free PMC article.
-
Turbocharging introgression breeding of perennial fruit crops: a case study on apple.Hortic Res. 2020 Apr 1;7:47. doi: 10.1038/s41438-020-0270-z. eCollection 2020. Hortic Res. 2020. PMID: 32257233 Free PMC article.
-
Fine mapping by composite genome-wide association analysis.Genet Res (Camb). 2017 Jun 6;99:e4. doi: 10.1017/S0016672317000027. Genet Res (Camb). 2017. PMID: 28583209 Free PMC article.
-
Including α s1 casein gene information in genomic evaluations of French dairy goats.Genet Sel Evol. 2016 Aug 4;48(1):54. doi: 10.1186/s12711-016-0233-x. Genet Sel Evol. 2016. PMID: 27491470 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

