A surrogate approach to study the evolution of noncoding DNA elements that organize eukaryotic genomes
- PMID: 19635763
- PMCID: PMC2877547
- DOI: 10.1093/jhered/esp063
A surrogate approach to study the evolution of noncoding DNA elements that organize eukaryotic genomes
Abstract
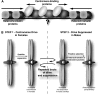
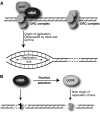
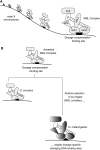
Comparative genomics provides a facile way to address issues of evolutionary constraint acting on different elements of the genome. However, several important DNA elements have not reaped the benefits of this new approach. Some have proved intractable to current day sequencing technology. These include centromeric and heterochromatic DNA, which are essential for chromosome segregation as well as gene regulation, but the highly repetitive nature of the DNA sequences in these regions make them difficult to assemble into longer contigs. Other sequences, like dosage compensation X chromosomal sites, origins of DNA replication, or heterochromatic sequences that encode piwi-associated RNAs, have proved difficult to study because they do not have recognizable DNA features that allow them to be described functionally or computationally. We have employed an alternate approach to the direct study of these DNA elements. By using proteins that specifically bind these noncoding DNAs as surrogates, we can indirectly assay the evolutionary constraints acting on these important DNA elements. We review the impact that such "surrogate strategies" have had on our understanding of the evolutionary constraints shaping centromeres, origins of DNA replication, and dosage compensation X chromosomal sites. These have begun to reveal that in contrast to the view that such structural DNA elements are either highly constrained (under purifying selection) or free to drift (under neutral evolution), some of them may instead be shaped by adaptive evolution and genetic conflicts (these are not mutually exclusive). These insights also help to explain why the same elements (e.g., centromeres and replication origins), which are so complex in some eukaryotic genomes, can be simple and well defined in other where similar conflicts do not exist.
Figures

References
-
- Akhtar A, Becker PB. Activation of transcription through histone H4 acetylation by MOF, an acetyltransferase essential for dosage compensation in Drosophila. Mol Cell. 2000;5:367–375. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

