Chromosome aberrations resulting from double-strand DNA breaks at a naturally occurring yeast fragile site composed of inverted ty elements are independent of Mre11p and Sae2p
- PMID: 19635935
- PMCID: PMC2766307
- DOI: 10.1534/genetics.109.106385
Chromosome aberrations resulting from double-strand DNA breaks at a naturally occurring yeast fragile site composed of inverted ty elements are independent of Mre11p and Sae2p
Abstract
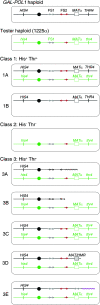
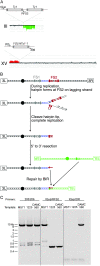
Genetic instability at palindromes and spaced inverted repeats (IRs) leads to chromosome rearrangements. Perfect palindromes and IRs with short spacers can extrude as cruciforms or fold into hairpins on the lagging strand during replication. Cruciform resolution produces double-strand breaks (DSBs) with hairpin-capped ends, and Mre11p and Sae2p are required to cleave the hairpin tips to facilitate homologous recombination. Fragile site 2 (FS2) is a naturally occurring IR in Saccharomyces cerevisiae composed of a pair of Ty1 elements separated by approximately 280 bp. Our results suggest that FS2 forms a hairpin, rather than a cruciform, during replication in cells with low levels of DNA polymerase. Cleavage of this hairpin results in a recombinogenic DSB. We show that DSB formation at FS2 does not require Mre11p, Sae2p, Rad1p, Slx4p, Pso2p, Exo1p, Mus81p, Yen1p, or Rad27p. Also, repair of DSBs by homologous recombination is efficient in mre11 and sae2 mutants. Homologous recombination is impaired at FS2 in rad52 mutants and most aberrations reflect either joining of two broken chromosomes in a "half crossover" or telomere capping of the break. In support of hairpin formation precipitating DSBs at FS2, two telomere-capped deletions had a breakpoint near the center of the IR. In summary, Mre11p and Sae2p are not required for DSB formation at FS2 or the subsequent repair of these DSBs.
Figures




References
-
- Allers, T., and D. R. Leach, 1995. DNA palindromes adopt a methylation-resistant conformation that is consistent with DNA cruciform or hairpin formation in vivo. J. Mol. Biol. 252: 70–85. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

