High-throughput molecular analysis in lung cancer: insights into biology and potential clinical applications
- PMID: 19648524
- PMCID: PMC4648268
- DOI: 10.1183/09031936.00042409
High-throughput molecular analysis in lung cancer: insights into biology and potential clinical applications
Abstract
During the last decade, high-throughput technologies including genomic, epigenomic, transcriptomic and proteomic have been applied to further our understanding of the molecular pathogenesis of this heterogeneous disease, and to develop strategies that aim to improve the management of patients with lung cancer. Ultimately, these approaches should lead to sensitive, specific and noninvasive methods for early diagnosis, and facilitate the prediction of response to therapy and outcome, as well as the identification of potential novel therapeutic targets. Genomic studies were the first to move this field forward by providing novel insights into the molecular biology of lung cancer and by generating candidate biomarkers of disease progression. Lung carcinogenesis is driven by genetic and epigenetic alterations that cause aberrant gene function; however, the challenge remains to pinpoint the key regulatory control mechanisms and to distinguish driver from passenger alterations that may have a small but additive effect on cancer development. Epigenetic regulation by DNA methylation and histone modifications modulate chromatin structure and, in turn, either activate or silence gene expression. Proteomic approaches critically complement these molecular studies, as the phenotype of a cancer cell is determined by proteins and cannot be predicted by genomics or transcriptomics alone. The present article focuses on the technological platforms available and some proposed clinical applications. We illustrate herein how the "-omics" have revolutionised our approach to lung cancer biology and hold promise for personalised management of lung cancer.
Conflict of interest statement
A statement of interest for R.K. Thomas can be found at
Figures



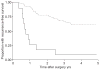

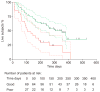
Similar articles
-
Epigenomic targets for the treatment of respiratory disease.Expert Opin Ther Targets. 2009 Jun;13(6):625-40. doi: 10.1517/14728220902926119. Expert Opin Ther Targets. 2009. PMID: 19409032 Review.
-
Epigenetic biomarkers in lung cancer.Cancer Lett. 2014 Jan 28;342(2):200-12. doi: 10.1016/j.canlet.2012.04.018. Epub 2012 Apr 27. Cancer Lett. 2014. PMID: 22546286 Review.
-
Identification of Epigenetic Biomarkers of Lung Adenocarcinoma through Multi-Omics Data Analysis.PLoS One. 2016 Apr 4;11(4):e0152918. doi: 10.1371/journal.pone.0152918. eCollection 2016. PLoS One. 2016. PMID: 27042856 Free PMC article.
-
Epigenetics in lung cancer diagnosis and therapy.Cancer Metastasis Rev. 2015 Jun;34(2):229-41. doi: 10.1007/s10555-015-9563-3. Cancer Metastasis Rev. 2015. PMID: 25939322 Review.
-
Lung cancer epigenetics: From knowledge to applications.Semin Cancer Biol. 2018 Aug;51:116-128. doi: 10.1016/j.semcancer.2017.09.005. Epub 2017 Sep 14. Semin Cancer Biol. 2018. PMID: 28919484 Review.
Cited by
-
Extracellular matrix gene expression profiling using microfluidics for colorectal carcinoma stratification.Biomicrofluidics. 2016 Oct 31;10(5):054124. doi: 10.1063/1.4966245. eCollection 2016 Sep. Biomicrofluidics. 2016. PMID: 27822332 Free PMC article.
-
Proteomic approaches in biomarker discovery: new perspectives in cancer diagnostics.ScientificWorldJournal. 2014 Jan 14;2014:260348. doi: 10.1155/2014/260348. eCollection 2014. ScientificWorldJournal. 2014. PMID: 24550697 Free PMC article. Review.
-
Transcriptomic signals in blood prior to lung cancer focusing on time to diagnosis and metastasis.Sci Rep. 2021 Apr 1;11(1):7406. doi: 10.1038/s41598-021-86879-8. Sci Rep. 2021. PMID: 33795786 Free PMC article.
-
Epidemiology of lung cancer: Diagnosis and management of lung cancer, 3rd ed: American College of Chest Physicians evidence-based clinical practice guidelines.Chest. 2013 May;143(5 Suppl):e1S-e29S. doi: 10.1378/chest.12-2345. Chest. 2013. PMID: 23649439 Free PMC article. Review.
-
An immune-related nomogram model that predicts the overall survival of patients with lung adenocarcinoma.BMC Pulm Med. 2022 Mar 30;22(1):114. doi: 10.1186/s12890-022-01902-6. BMC Pulm Med. 2022. PMID: 35354459 Free PMC article.
References
-
- Parkin DM, Bray F, Ferlay J, et al. Global cancer statistics, 2002. CA Cancer J Clin. 2005;55:74–108. - PubMed
-
- Giaccone G. Clinical impact of novel treatment strategies. Oncogene. 2002;21:6970–6981. - PubMed
-
- Winton T, Livingston R, Johnson D, et al. Vinorelbine plus cisplatin vs observation in resected non-small-cell lung cancer. N Engl J Med. 2005;352:2589–2597. - PubMed
-
- Pisters KM, Le Chevalier T. Adjuvant chemotherapy in completely resected non-small-cell lung cancer. J Clin Oncol. 2005;23:3270–3278. - PubMed
-
- Chanin TD, Merrick DT, Franklin WA, et al. Recent developments in biomarkers for the early detection of lung cancer: perspectives based on publications 2003 to present. Curr Opin Pulm Med. 2004;10:242–247. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
