Evolutionary capture of viral and plasmid DNA by yeast nuclear chromosomes
- PMID: 19666779
- PMCID: PMC2756859
- DOI: 10.1128/EC.00110-09
Evolutionary capture of viral and plasmid DNA by yeast nuclear chromosomes
Abstract
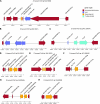
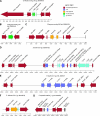
A 10-kb region of the nuclear genome of the yeast Vanderwaltozyma polyspora contains an unusual cluster of five pseudogenes homologous to five different genes from yeast killer viruses, killer plasmids, the 2microm plasmid, and a Penicillium virus. By further database searches, we show that this phenomenon is not unique to V. polyspora but that about 40% of the sequenced genomes of Saccharomycotina species contain integrated copies of genes from DNA plasmids or RNA viruses. We propose the name NUPAVs (nuclear sequences of plasmid and viral origin) for these objects, by analogy to NUMTs (nuclear copies of mitochondrial DNA) and NUPTs (nuclear copies of plastid DNA, in plants) of organellar origin. Although most of the NUPAVs are pseudogenes, one intact and active gene that was formed in this way is the KHS1 chromosomal killer locus of Saccharomyces cerevisiae. We show that KHS1 is a NUPAV related to M2 killer virus double-stranded RNA. Many NUPAVs are located beside tRNA genes, and some contain sequences from a mixture of different extrachromosomal sources. We propose that NUPAVs are sequences that were captured by the nuclear genome during the repair of double-strand breaks that occurred during evolution and that some of their properties may be explained by repeated breakage at fragile chromosomal sites.
Figures
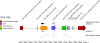

References
-
- Baeza, M. E., M. A. Sanhueza, and V. H. Cifuentes. 2008. Occurrence of killer yeast strains in industrial and clinical yeast isolates. Biol. Res. 41:173-182. - PubMed
-
- Blaisonneau, J., F. Sor, G. Cheret, D. Yarrow, and H. Fukuhara. 1997. A circular plasmid from the yeast Torulaspora delbrueckii. Plasmid 38:202-209. - PubMed
-
- Bostian, K. A., Q. Elliott, H. Bussey, V. Burn, A. Smith, and D. J. Tipper. 1984. Sequence of the preprotoxin dsRNA gene of type I killer yeast: multiple processing events produce a two-component toxin. Cell 36:741-751. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

