Structural studies of soybean calmodulin isoform 4 bound to the calmodulin-binding domain of tobacco mitogen-activated protein kinase phosphatase-1 provide insights into a sequential target binding mode
- PMID: 19667066
- PMCID: PMC2788880
- DOI: 10.1074/jbc.M109.025080
Structural studies of soybean calmodulin isoform 4 bound to the calmodulin-binding domain of tobacco mitogen-activated protein kinase phosphatase-1 provide insights into a sequential target binding mode
Abstract
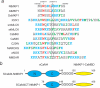
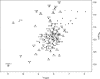
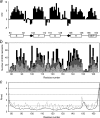
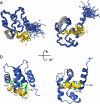
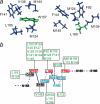
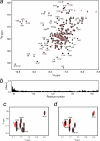
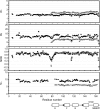
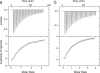
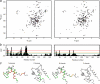
The calcium regulatory protein calmodulin (CaM) binds in a calcium-dependent manner to numerous target proteins. The calmodulin-binding domain (CaMBD) region of Nicotiana tabacum MAPK phosphatase has an amino acid sequence that does not resemble the CaMBD of any other known Ca(2+)-CaM-binding proteins. Using a unique fusion protein strategy, we have been able to obtain a high resolution solution structure of the complex of soybean Ca(2+)-CaM4 (SCaM4) and this CaMBD. Complete isotope labeling of both parts of the complex in the fusion protein greatly facilitated the structure determination by NMR. The 12-residue CaMBD region was found to bind exclusively to the C-lobe of SCaM4. A specific Trp and Leu side chain are utilized to facilitate strong binding through a novel "double anchor" motif. Moreover, the orientation of the helical peptide on the surface of Ca(2+)-SCaM4 is distinct from other known complexes. The N-lobe of Ca(2+)-SCaM4 in the complex remains free for additional interactions and could possibly act as a calcium-dependent adapter protein. Signaling through the MAPK pathway and increases in intracellular Ca(2+) are both hallmarks of the plant stress response, and our data support the notion that coordination of these responses may occur through the formation of a unique CaM-MAPK phosphatase multiprotein complex.
Figures
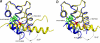
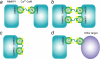
Similar articles
-
Regulation of MAPK phosphatase 1 (AtMKP1) by calmodulin in Arabidopsis.J Biol Chem. 2008 Aug 29;283(35):23581-8. doi: 10.1074/jbc.M801549200. Epub 2008 Jun 25. J Biol Chem. 2008. PMID: 18579522 Free PMC article.
-
The solution structures of two soybean calmodulin isoforms provide a structural basis for their selective target activation properties.J Biol Chem. 2008 May 23;283(21):14619-28. doi: 10.1074/jbc.M801398200. Epub 2008 Mar 17. J Biol Chem. 2008. PMID: 18347016
-
Calcium-dependent and -independent binding of soybean calmodulin isoforms to the calmodulin binding domain of tobacco MAPK phosphatase-1.J Biol Chem. 2007 Mar 2;282(9):6031-42. doi: 10.1074/jbc.M608970200. Epub 2007 Jan 3. J Biol Chem. 2007. PMID: 17202149
-
Calmodulin's flexibility allows for promiscuity in its interactions with target proteins and peptides.Mol Biotechnol. 2004 May;27(1):33-57. doi: 10.1385/MB:27:1:33. Mol Biotechnol. 2004. PMID: 15122046 Review.
-
A Non-Canonical Calmodulin Target Motif Comprising a Polybasic Region and Lipidated Terminal Residue Regulates Localization.Int J Mol Sci. 2020 Apr 15;21(8):2751. doi: 10.3390/ijms21082751. Int J Mol Sci. 2020. PMID: 32326637 Free PMC article. Review.
Cited by
-
Structural characterization of the interaction of human lactoferrin with calmodulin.PLoS One. 2012;7(12):e51026. doi: 10.1371/journal.pone.0051026. Epub 2012 Dec 6. PLoS One. 2012. PMID: 23236421 Free PMC article.
-
Unraveling the Potential of γ-Aminobutyric Acid: Insights into Its Biosynthesis and Biotechnological Applications.Nutrients. 2024 Aug 19;16(16):2760. doi: 10.3390/nu16162760. Nutrients. 2024. PMID: 39203897 Free PMC article. Review.
-
Structural insights into calmodulin-regulated L-selectin ectodomain shedding.J Biol Chem. 2012 Aug 3;287(32):26513-27. doi: 10.1074/jbc.M112.373373. Epub 2012 Jun 18. J Biol Chem. 2012. PMID: 22711531 Free PMC article.
-
Design of high-affinity S100-target hybrid proteins.Protein Sci. 2009 Dec;18(12):2528-36. doi: 10.1002/pro.267. Protein Sci. 2009. PMID: 19827097 Free PMC article.
-
Structural basis for the regulation of L-type voltage-gated calcium channels: interactions between the N-terminal cytoplasmic domain and Ca(2+)-calmodulin.Front Mol Neurosci. 2012 Apr 12;5:38. doi: 10.3389/fnmol.2012.00038. eCollection 2012. Front Mol Neurosci. 2012. PMID: 22518098 Free PMC article.
References
-
- Chin D., Means A. R. (2000) Trends Cell Biol. 10, 322–328 - PubMed
-
- Hoeflich K. P., Ikura M. (2002) Cell 108, 739–742 - PubMed
-
- Yamauchi E., Nakatsu T., Matsubara M., Kato H., Taniguchi H. (2003) Nat. Struct. Biol. 10, 226–231 - PubMed
-
- Maximciuc A. A., Putkey J. A., Shamoo Y., Mackenzie K. R. (2006) Structure 14, 1547–1556 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Miscellaneous

