The evolution of the major hepatitis C genotypes correlates with clinical response to interferon therapy
- PMID: 19668364
- PMCID: PMC2719056
- DOI: 10.1371/journal.pone.0006579
The evolution of the major hepatitis C genotypes correlates with clinical response to interferon therapy
Abstract
Background: Patients chronically infected with hepatitis C virus (HCV) require significantly different durations of therapy and achieve substantially different sustained virologic response rates to interferon-based therapies, depending on the HCV genotype with which they are infected. There currently exists no systematic framework that explains these genotype-specific response rates. Since humans are the only known natural hosts for HCV-a virus that is at least hundreds of years old-one possibility is that over the time frame of this relationship, HCV accumulated adaptive mutations that confer increasing resistance to the human immune system. Given that interferon therapy functions by triggering an immune response, we hypothesized that clinical response rates are a reflection of viral evolutionary adaptations to the immune system.
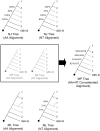
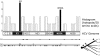
Methods and findings: We have performed the first phylogenetic analysis to include all available full-length HCV genomic sequences (n = 345). This resulted in a new cladogram of HCV. This tree establishes for the first time the relative evolutionary ages of the major HCV genotypes. The outcome data from prospective clinical trials that studied interferon and ribavirin therapy was then mapped onto this new tree. This mapping revealed a correlation between genotype-specific responses to therapy and respective genotype age. This correlation allows us to predict that genotypes 5 and 6, for which there currently are no published prospective trials, will likely have intermediate response rates, similar to genotype 3. Ancestral protein sequence reconstruction was also performed, which identified the HCV proteins E2 and NS5A as potential determinants of genotype-specific clinical outcome. Biochemical studies have independently identified these same two proteins as having genotype-specific abilities to inhibit the innate immune factor double-stranded RNA-dependent protein kinase (PKR).
Conclusion: An evolutionary analysis of all available HCV genomes supports the hypothesis that immune selection was a significant driving force in the divergence of the major HCV genotypes and that viral factors that acquired the ability to inhibit the immune response may play a role in determining genotype-specific response rates to interferon therapy.
Conflict of interest statement
Figures



Similar articles
-
Analysis of sequences of hepatitis C virus NS5A genotype 1 in HIV-coinfected patients with a null response to nitazoxanide or peg-interferon plus ribavirin.Arch Virol. 2013 Sep;158(9):1907-15. doi: 10.1007/s00705-013-1687-6. Epub 2013 Apr 4. Arch Virol. 2013. PMID: 23553458
-
Analysis of the PKR-eIF2alpha phosphorylation homology domain (PePHD) of hepatitis C virus genotype 1 in HIV-coinfected patients by ultra-deep pyrosequencing and its relationship to responses to pegylated interferon-ribavirin treatment.Arch Virol. 2012 Apr;157(4):703-11. doi: 10.1007/s00705-012-1230-1. Epub 2012 Jan 22. Arch Virol. 2012. PMID: 22270759
-
Genetic variability of hepatitis C virus in chronically infected patients with viral breakthrough during interferon-ribavirin therapy.J Med Virol. 2004 Sep;74(1):41-53. doi: 10.1002/jmv.20144. J Med Virol. 2004. PMID: 15258967
-
Understanding the molecular mechanism(s) of hepatitis C virus (HCV) induced interferon resistance.Infect Genet Evol. 2013 Oct;19:113-9. doi: 10.1016/j.meegid.2013.06.025. Epub 2013 Jul 5. Infect Genet Evol. 2013. PMID: 23831932 Review.
-
Viral, host and interferon-related factors modulating the effect of interferon therapy for hepatitis C virus infection.J Viral Hepat. 2001 Jan;8(1):1-18. doi: 10.1046/j.1365-2893.2001.00253.x. J Viral Hepat. 2001. PMID: 11155147 Review.
Cited by
-
Global virus outbreaks: Interferons as 1st responders.Semin Immunol. 2019 Jun;43:101300. doi: 10.1016/j.smim.2019.101300. Semin Immunol. 2019. PMID: 31771760 Free PMC article. Review.
-
Lipids and HCV.Semin Immunopathol. 2013 Jan;35(1):87-100. doi: 10.1007/s00281-012-0356-2. Epub 2012 Oct 31. Semin Immunopathol. 2013. PMID: 23111699 Review.
-
Regulation of hepatic innate immunity by hepatitis C virus.Nat Med. 2013 Jul;19(7):879-88. doi: 10.1038/nm.3253. Nat Med. 2013. PMID: 23836238 Free PMC article. Review.
-
Computational analysis of naturally occurring resistance-associated substitutions in genes NS3, NS5A, and NS5B among 86 subtypes of hepatitis C virus worldwide.Infect Drug Resist. 2019 Sep 19;12:2987-3015. doi: 10.2147/IDR.S218584. eCollection 2019. Infect Drug Resist. 2019. PMID: 31571951 Free PMC article.
-
Comparison of the Mechanisms of Drug Resistance among HIV, Hepatitis B, and Hepatitis C.Viruses. 2010 Dec 1;2(12):2696-739. doi: 10.3390/v2122696. Viruses. 2010. PMID: 21243082 Free PMC article.
References
-
- National Institutes of Health Consensus Development Conference Statement: Management of hepatitis C 2002 (June 10–12, 2002). Gastroenterology. 2002;123:2082–2099. - PubMed
-
- Verna EC, Brown RS,, Jr. Hepatitis C virus and liver transplantation. Clin Liver Dis. 2006;10:919–940. - PubMed
-
- Hofmann WP, Herrmann E, Sarrazin C, Zeuzem S. Ribavirin mode of action in chronic hepatitis C: from clinical use back to molecular mechanisms. Liver Int. 2008;28:1332–1343. - PubMed
-
- Feld JJ, Hoofnagle JH. Mechanism of action of interferon and ribavirin in treatment of hepatitis C. Nature. 2005;436:967–972. - PubMed
-
- Mangia A, Santoro R, Minerva N, Ricci GL, Carretta V, et al. Peginterferon alfa-2b and ribavirin for 12 vs. 24 weeks in HCV genotype 2 or 3. N Engl J Med. 2005;352:2609–2617. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

