A genome-wide compilation of the two-component systems in Lotus japonicus
- PMID: 19675111
- PMCID: PMC2725789
- DOI: 10.1093/dnares/dsp012
A genome-wide compilation of the two-component systems in Lotus japonicus
Abstract
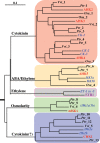
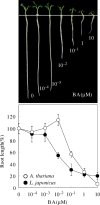
The two-component systems (TCS), or histidine-to-aspartate phosphorelays, are evolutionarily conserved common signal transduction mechanisms that are implicated in a wide variety of cellular responses to environmental stimuli in both prokaryotes and eukaryotes including plants. Among higher plants, legumes including Lotus japonicus have a unique ability to engage in beneficial symbiosis with nitrogen-fixing bacteria. We previously presented a genome-wide compiled list of TCS-associated components of Mesorhizobium loti, which is a symbiont specific to L. japonicus (Hagiwara et al. 2004, DNA Res., 11, 57-65). To gain both general and specific insights into TCS of this currently attractive model legume, here we compiled TCS-associated components as many as possible from a genome-wide viewpoint by taking advantage that the efforts of whole genome sequencing of L. japonicus are almost at final stage. In the current database (http://www.kazusa.or.jp/lotus/index.html), it was found that L. japonicus has, at least, 14 genes each encoding a histidine kinase, 7 histidine-containing phosphotransmitter-related genes, 7 type-A response regulator (RR)-related genes, 11 type-B RR-related genes, and also 5 circadian clock-associated pseudo-RR genes. These results suggested that most of the L. japonicus TCS-associated genes have already been uncovered in this genome-wide analysis, if not all. Here, characteristics of these TCS-associated components of L. japonicus were inspected, one by one, in comparison with those of Arabidopsis thaliana. In addition, some critical experiments were also done to gain further insights into the functions of L. japonicus TCS-associated genes with special reference to cytokinin-mediated signal transduction and circadian clock.
Figures





References
-
- Mizuno T. His-Asp phosphotrasnsfer signal transduction. J. Biochem. (Tokyo) 1998;123:555–63. - PubMed
-
- Stock A.M., Robinson V.L., Goudreau P.N. Two-component signal transduction. Annu. Rev. Biochem. 2000;69:183–215. - PubMed
-
- Mizuno T. Plant response regulators implicated in signal transduction and circadian rhythm. Curr. Opin. Plant Biol. 2004;7:499–505. - PubMed
-
- Mizuno T. Two-component phosphorelay signal transduction systems in plants: from hormone response to circadian rhythms. Biosci. Biotechnol, Biochem. 2005;69:2263–76. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

