Regulation of sulfotransferase and UDP-glucuronosyltransferase gene expression by the PPARs
- PMID: 19680455
- PMCID: PMC2724710
- DOI: 10.1155/2009/728941
Regulation of sulfotransferase and UDP-glucuronosyltransferase gene expression by the PPARs
Abstract
During phase II metabolism, a substrate is rendered more hydrophilic through the covalent attachment of an endogenous molecule. The cytosolic sulfotransferase (SULT) and UDP-glucuronosyltransferase (UGT) families of enzymes account for the majority of phase II metabolism in humans and animals. In general, phase II metabolism is considered to be a detoxication process, as sulfate and glucuronide conjugates are more amenable to excretion and elimination than are the parent substrates. However, certain products of phase II metabolism (e.g., unstable sulfate conjugates) are genotoxic. Members of the nuclear receptor superfamily are particularly important regulators of SULT and UGT gene transcription. In metabolically active tissues, increasing evidence supports a major role for lipid-sensing transcription factors, such as peroxisome proliferator-activated receptors (PPARs), in the regulation of rodent and human SULT and UGT gene expression. This review summarizes current information regarding the regulation of these two major classes of phase II metabolizing enzyme by PPARs.
Figures
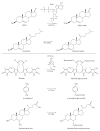


Similar articles
-
Multiple UDP-Glucuronosyltransferase and Sulfotransferase Enzymes are Responsible for the Metabolism of Verproside in Human Liver Preparations.Molecules. 2017 Apr 22;22(4):670. doi: 10.3390/molecules22040670. Molecules. 2017. PMID: 28441724 Free PMC article.
-
Peroxisome proliferator-activated receptor alpha induces hepatic expression of the human bile acid glucuronidating UDP-glucuronosyltransferase 2B4 enzyme.J Biol Chem. 2003 Aug 29;278(35):32852-60. doi: 10.1074/jbc.M305361200. Epub 2003 Jun 16. J Biol Chem. 2003. PMID: 12810707
-
Identification of Catalposide Metabolites in Human Liver and Intestinal Preparations and Characterization of the Relevant Sulfotransferase, UDP-glucuronosyltransferase, and Carboxylesterase Enzymes.Pharmaceutics. 2019 Jul 22;11(7):355. doi: 10.3390/pharmaceutics11070355. Pharmaceutics. 2019. PMID: 31336576 Free PMC article.
-
Peroxisome proliferators and peroxisome proliferator activated receptors (PPARs) as regulators of lipid metabolism.Biochimie. 1997 Feb-Mar;79(2-3):81-94. doi: 10.1016/s0300-9084(97)81496-4. Biochimie. 1997. PMID: 9209701 Review.
-
The UDP-glucuronosyltransferases: their role in drug metabolism and detoxification.Int J Biochem Cell Biol. 2013 Jun;45(6):1121-32. doi: 10.1016/j.biocel.2013.02.019. Epub 2013 Mar 7. Int J Biochem Cell Biol. 2013. PMID: 23500526 Review.
Cited by
-
Updated perspectives on the cytosolic sulfotransferases (SULTs) and SULT-mediated sulfation.Biosci Biotechnol Biochem. 2017 Jan;81(1):63-72. doi: 10.1080/09168451.2016.1222266. Epub 2016 Sep 21. Biosci Biotechnol Biochem. 2017. PMID: 27649811 Free PMC article. Review.
-
Cytosolic sulfotransferases in endocrine disruption.Essays Biochem. 2024 Dec 4;68(4):541-553. doi: 10.1042/EBC20230101. Essays Biochem. 2024. PMID: 38699885 Free PMC article. Review.
-
PPARα: A Master Regulator of Bilirubin Homeostasis.PPAR Res. 2014;2014:747014. doi: 10.1155/2014/747014. Epub 2014 Jul 23. PPAR Res. 2014. PMID: 25147562 Free PMC article.
-
Current View on PPAR-α and Its Relation to Neurosteroids in Alzheimer's Disease and Other Neuropsychiatric Disorders: Promising Targets in a Therapeutic Strategy.Int J Mol Sci. 2024 Jun 28;25(13):7106. doi: 10.3390/ijms25137106. Int J Mol Sci. 2024. PMID: 39000217 Free PMC article. Review.
-
Changes in the Metabolome in Response to Low-Dose Exposure to Environmental Chemicals Used in Personal Care Products during Different Windows of Susceptibility.PLoS One. 2016 Jul 28;11(7):e0159919. doi: 10.1371/journal.pone.0159919. eCollection 2016. PLoS One. 2016. PMID: 27467775 Free PMC article.
References
-
- Robinson E, Grieve DJ. Significance of peroxisome proliferator-activated receptors in the cardiovascular system in health and disease. Pharmacology & Therapeutics. 2009;122(3):246–263. - PubMed
-
- Töröcsik D, Szanto A, Nagy L. Oxysterol signaling links cholesterol metabolism and inflammation via the liver X receptor in macrophages. Molecular Aspects of Medicine. 2009;30(3):134–152. - PubMed
-
- Chapman E, Best MD, Hanson SR, Wong C-H. Sulfotransferases: structure, mechanism, biological activity, inhibition, and synthetic utility. Angewandte Chemie International Edition. 2004;43(27):3526–3548. - PubMed
-
- Strott CA. Sulfonation and molecular action. Endocrine Reviews. 2002;23(5):703–732. - PubMed
-
- Dooley TP, Haldeman-Cahill R, Joiner J, Wilborn TW. Expression profiling of human sulfotransferase and sulfatase gene superfamilies in epithelial tissues and cultured cells. Biochemical and Biophysical Research Communications. 2000;277(1):236–245. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

