Genome-wide mRNA expression analysis of hepatic adaptation to high-fat diets reveals switch from an inflammatory to steatotic transcriptional program
- PMID: 19680557
- PMCID: PMC2722023
- DOI: 10.1371/journal.pone.0006646
Genome-wide mRNA expression analysis of hepatic adaptation to high-fat diets reveals switch from an inflammatory to steatotic transcriptional program
Abstract
Background: Excessive exposure to dietary fats is an important factor in the initiation of obesity and metabolic syndrome associated pathologies. The cellular processes associated with the onset and progression of diet-induced metabolic syndrome are insufficiently understood.
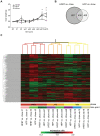
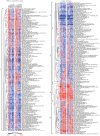
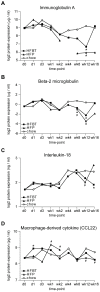
Principal findings: To identify the mechanisms underlying the pathological changes associated with short and long-term exposure to excess dietary fat, hepatic gene expression of ApoE3Leiden mice fed chow and two types of high-fat (HF) diets was monitored using microarrays during a 16-week period. A functional characterization of 1663 HF-responsive genes reveals perturbations in lipid, cholesterol and oxidative metabolism, immune and inflammatory responses and stress-related pathways. The major changes in gene expression take place during the early (day 3) and late (week 12) phases of HF feeding. This is also associated with characteristic opposite regulation of many HF-affected pathways between these two phases. The most prominent switch occurs in the expression of inflammatory/immune pathways (early activation, late repression) and lipogenic/adipogenic pathways (early repression, late activation). Transcriptional network analysis identifies NF-kappaB, NEMO, Akt, PPARgamma and SREBP1 as the key controllers of these processes and suggests that direct regulatory interactions between these factors may govern the transition from early (stressed, inflammatory) to late (pathological, steatotic) hepatic adaptation to HF feeding. This transition observed by hepatic gene expression analysis is confirmed by expression of inflammatory proteins in plasma and the late increase in hepatic triglyceride content. In addition, the genes most predictive of fat accumulation in liver during 16-week high-fat feeding period are uncovered by regression analysis of hepatic gene expression and triglyceride levels.
Conclusions: The transition from an inflammatory to a steatotic transcriptional program, possibly driven by the reciprocal activation of NF-kappaB and PPARgamma regulators, emerges as the principal signature of the hepatic adaptation to excess dietary fat. These findings may be of essential interest for devising new strategies aiming to prevent the progression of high-fat diet induced pathologies.
Conflict of interest statement
Figures






Similar articles
-
Dietary modification dampens liver inflammation and fibrosis in obesity-related fatty liver disease.Obesity (Silver Spring). 2013 Jun;21(6):1189-99. doi: 10.1002/oby.20123. Epub 2013 May 10. Obesity (Silver Spring). 2013. PMID: 23666886
-
Altered expression of transcription factors and genes regulating lipogenesis in liver and adipose tissue of mice with high fat diet-induced obesity and nonalcoholic fatty liver disease.Eur J Gastroenterol Hepatol. 2008 Sep;20(9):843-54. doi: 10.1097/MEG.0b013e3282f9b203. Eur J Gastroenterol Hepatol. 2008. PMID: 18794597
-
High-protein diets prevent steatosis and induce hepatic accumulation of monomethyl branched-chain fatty acids.J Nutr Biochem. 2014 Dec;25(12):1263-74. doi: 10.1016/j.jnutbio.2014.07.005. Epub 2014 Sep 16. J Nutr Biochem. 2014. PMID: 25287814
-
Regulation of the apolipoprotein E by dietary lipids occurs by transcriptional and post-transcriptional mechanisms.Mol Cell Biochem. 1996 Feb 23;155(2):153-62. doi: 10.1007/BF00229312. Mol Cell Biochem. 1996. PMID: 8700160
-
Genetics in Light of Transcriptional Adaptation.Trends Genet. 2020 Dec;36(12):926-935. doi: 10.1016/j.tig.2020.08.008. Epub 2020 Sep 11. Trends Genet. 2020. PMID: 32928563 Free PMC article. Review.
Cited by
-
The effect of dietary fat intake on hepatic gene expression in LG/J AND SM/J mice.BMC Genomics. 2014 Feb 5;15:99. doi: 10.1186/1471-2164-15-99. BMC Genomics. 2014. PMID: 24499025 Free PMC article.
-
The development of diet-induced obesity and glucose intolerance in C57BL/6 mice on a high-fat diet consists of distinct phases.PLoS One. 2014 Aug 29;9(8):e106159. doi: 10.1371/journal.pone.0106159. eCollection 2014. PLoS One. 2014. PMID: 25170916 Free PMC article.
-
Oligonucleotide microarray analysis of dietary-induced hyperlipidemia gene expression profiles in miniature pigs.PLoS One. 2012;7(5):e37581. doi: 10.1371/journal.pone.0037581. Epub 2012 May 25. PLoS One. 2012. PMID: 22662175 Free PMC article.
-
Antiobesity Effects of an Edible Halophyte Nitraria retusa Forssk in 3T3-L1 Preadipocyte Differentiation and in C57B6J/L Mice Fed a High Fat Diet-Induced Obesity.Evid Based Complement Alternat Med. 2013;2013:368658. doi: 10.1155/2013/368658. Epub 2013 Dec 3. Evid Based Complement Alternat Med. 2013. PMID: 24367387 Free PMC article.
-
A Moderate Intake of Beer Improves Metabolic Dysfunction-Associated Steatotic Liver Disease (MASLD) in a High-Fat Diet (HFD)-Induced Mouse Model.Molecules. 2024 Dec 17;29(24):5954. doi: 10.3390/molecules29245954. Molecules. 2024. PMID: 39770043 Free PMC article.
References
-
- Kopelman PG. Obesity as a medical problem. Nature. 2000;404:635–643. - PubMed
-
- Ogden CL, Yanovski SZ, Carroll MD, Flegal KM. The epidemiology of obesity. Gastroenterology. 2007;132:2087–2102. - PubMed
-
- Lissner L, Heitmann BL. Dietary fat and obesity: evidence from epidemiology. Eur J Clin Nutr. 1995;49:79–90. - PubMed
-
- Buettner R, Scholmerich J, Bollheimer LC. High-fat diets: modeling the metabolic disorders of human obesity in rodents. Obesity (Silver Spring) 2007;15:798–808. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

