What can you do with 0.1x genome coverage? A case study based on a genome survey of the scuttle fly Megaselia scalaris (Phoridae)
- PMID: 19689807
- PMCID: PMC2735751
- DOI: 10.1186/1471-2164-10-382
What can you do with 0.1x genome coverage? A case study based on a genome survey of the scuttle fly Megaselia scalaris (Phoridae)
Abstract
Background: The declining cost of DNA sequencing is making genome sequencing a feasible option for more organisms, including many of interest to ecologists and evolutionary biologists. While obtaining high-depth, completely assembled genome sequences for most non-model organisms remains challenging, low-coverage genome survey sequences (GSS) can provide a wealth of biologically useful information at low cost. Here, using a random pyrosequencing approach, we sequence the genome of the scuttle fly Megaselia scalaris and evaluate the utility of our low-coverage GSS approach.
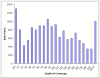
Results: Random pyrosequencing of the M. scalaris genome provided a depth of coverage (0.05x0.1x) much lower than typical GSS studies. We demonstrate that, even with extremely low-coverage sequencing, bioinformatics approaches can yield extensive information about functional and repetitive elements. We also use our GSS data to develop genomic resources such as a nearly complete mitochondrial genome sequence and microsatellite markers for M. scalaris.
Conclusion: We conclude that low-coverage genome surveys are effective at generating useful information about organisms currently lacking genomic sequence data.
Figures

Similar articles
-
The complete mitochondrial genome of the scuttle fly, Megaselia scalaris (Diptera: Phoridae).Mitochondrial DNA A DNA Mapp Seq Anal. 2016;27(1):182-4. doi: 10.3109/19401736.2013.879651. Epub 2014 Feb 3. Mitochondrial DNA A DNA Mapp Seq Anal. 2016. PMID: 24491096
-
Morphological and Molecular Characteristic of Megaselia scalaris (Diptera: Phoridae) Larvae as the Cause of Urinary Myiasis.J Med Entomol. 2017 May 1;54(3):781-784. doi: 10.1093/jme/tjw204. J Med Entomol. 2017. PMID: 28399213
-
Scuttle fly Megaselia scalaris (Loew) (Diptera: Phoridae) endoparasitoid as a novel biocontrol agent against adult American cockroaches (Periplaneta americana).Sci Rep. 2024 Apr 29;14(1):9762. doi: 10.1038/s41598-024-59547-w. Sci Rep. 2024. PMID: 38684676 Free PMC article.
-
Natural history of the scuttle fly, Megaselia scalaris.Annu Rev Entomol. 2008;53:39-60. doi: 10.1146/annurev.ento.53.103106.093415. Annu Rev Entomol. 2008. PMID: 17622197 Review.
-
Helitrons shaping the genomic architecture of Drosophila: enrichment of DINE-TR1 in α- and β-heterochromatin, satellite DNA emergence, and piRNA expression.Chromosome Res. 2015 Sep;23(3):597-613. doi: 10.1007/s10577-015-9480-x. Chromosome Res. 2015. PMID: 26408292 Review.
Cited by
-
Characterization of Plastidial and Nuclear SSR Markers for Understanding Invasion Histories and Genetic Diversity of Schinus molle L.Biology (Basel). 2018 Aug 10;7(3):43. doi: 10.3390/biology7030043. Biology (Basel). 2018. PMID: 30103413 Free PMC article.
-
Mitochondrial genomes of two phlebotomine sand flies, Phlebotomus chinensis and Phlebotomus papatasi (Diptera: Nematocera), the first representatives from the family Psychodidae.Parasit Vectors. 2015 Sep 17;8:472. doi: 10.1186/s13071-015-1081-1. Parasit Vectors. 2015. PMID: 26381614 Free PMC article.
-
Building a model: developing genomic resources for common milkweed (Asclepias syriaca) with low coverage genome sequencing.BMC Genomics. 2011 May 4;12:211. doi: 10.1186/1471-2164-12-211. BMC Genomics. 2011. PMID: 21542930 Free PMC article.
-
The complete mitochondrial genomes of two freshwater snails provide new protein-coding gene rearrangement models and phylogenetic implications.Parasit Vectors. 2017 Jan 6;10(1):11. doi: 10.1186/s13071-016-1956-9. Parasit Vectors. 2017. Retraction in: Parasit Vectors. 2017 Jul 21;10(1):350. doi: 10.1186/s13071-017-2287-1. PMID: 28061879 Free PMC article. Retracted.
-
Integration of molecular functions at the ecosystemic level: breakthroughs and future goals of environmental genomics and post-genomics.Ecol Lett. 2010 Jun;13(6):776-91. doi: 10.1111/j.1461-0248.2010.01464.x. Epub 2010 Apr 21. Ecol Lett. 2010. PMID: 20426792 Free PMC article. Review.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

