Examining polyglutamine peptide length: a connection between collapsed conformations and increased aggregation
- PMID: 19699209
- PMCID: PMC2764006
- DOI: 10.1016/j.jmb.2009.08.034
Examining polyglutamine peptide length: a connection between collapsed conformations and increased aggregation
Abstract
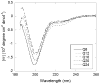
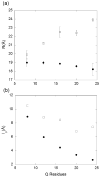
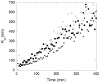
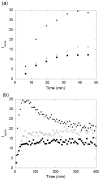
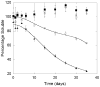
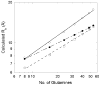
Abnormally expanded polyglutamine domains in proteins are associated with several neurodegenerative diseases, of which the best known is Huntington's. Expansion of the polyglutamine domain facilitates aggregation of the affected protein, and several studies directly link aggregation to neurotoxicity. The age of onset of disease is inversely correlated with the length of the polyglutamine domain; this correlation motivates an examination of the role of the length of the domain on aggregation. In this investigation, peptides containing 8 to 24 glutamines were synthesized, and their conformational and aggregation properties were examined. All peptides lacked secondary structure. Fluorescence resonance energy transfer studies revealed that the peptides became increasingly collapsed as the number of glutamine residues increased. The effective persistence length was estimated to decrease from approximately 11 to approximately 7 A as the number of glutamines increased from 8 to 24. A comparison of our data with theoretical results suggests that phosphate-buffered saline is a good solvent for Q8 and Q12, a theta solvent for Q16, and a poor solvent for Q20 and Q24. By dynamic light scattering, we observed that Q16, Q20, and Q24, but not Q8 or Q12, immediately formed soluble aggregates upon dilution into phosphate-buffered saline at 37 degrees C. Thus, Q16 stands at the transition point between good and poor solvent and between stable and aggregation-prone peptide. Examination of aggregates by transmission electron microscopy, along with kinetic assays for sedimentation, provided evidence indicating that soluble aggregates mature into sedimentable aggregates. Together, the data support a mechanism of aggregation in which monomer collapse is accompanied by formation of soluble oligomers; these soluble species lack regular secondary structure but appear morphologically similar to the sedimentable aggregates into which they eventually mature.
Figures


Similar articles
-
Elongation kinetics of polyglutamine peptide fibrils: a quartz crystal microbalance with dissipation study.J Mol Biol. 2012 Aug 10;421(2-3):329-47. doi: 10.1016/j.jmb.2012.03.017. Epub 2012 Mar 26. J Mol Biol. 2012. PMID: 22459263 Free PMC article.
-
Aggregation kinetics of interrupted polyglutamine peptides.J Mol Biol. 2011 Sep 23;412(3):505-19. doi: 10.1016/j.jmb.2011.07.003. Epub 2011 Jul 29. J Mol Biol. 2011. PMID: 21821045 Free PMC article.
-
Reconsidering the mechanism of polyglutamine peptide aggregation.Biochemistry. 2007 Nov 6;46(44):12810-20. doi: 10.1021/bi700806c. Epub 2007 Oct 11. Biochemistry. 2007. PMID: 17929830
-
Towards the treatment of polyglutamine diseases: the modulatory role of protein context.Curr Med Chem. 2010;17(27):3058-68. doi: 10.2174/092986710791959800. Curr Med Chem. 2010. PMID: 20629626 Review.
-
Pathogenesis of polyglutamine disorders: aggregation revisited.Hum Mol Genet. 2003 Oct 15;12 Spec No 2:R173-86. doi: 10.1093/hmg/ddg295. Hum Mol Genet. 2003. PMID: 14504263 Review.
Cited by
-
Evaluation of Nanoparticle Tracking for Characterization of Fibrillar Protein Aggregates.AIChE J. 2014 Apr 1;60(4):1236-1244. doi: 10.1002/aic.14349. AIChE J. 2014. PMID: 25843955 Free PMC article.
-
β-hairpin-mediated nucleation of polyglutamine amyloid formation.J Mol Biol. 2013 Apr 12;425(7):1183-97. doi: 10.1016/j.jmb.2013.01.016. Epub 2013 Jan 23. J Mol Biol. 2013. PMID: 23353826 Free PMC article.
-
Trinucleotide repeats: a structural perspective.Front Neurol. 2013 Jun 20;4:76. doi: 10.3389/fneur.2013.00076. eCollection 2013. Front Neurol. 2013. PMID: 23801983 Free PMC article.
-
Q&A: repeat-containing proteins.Nat Struct Mol Biol. 2015 Dec;22(12):943-5. doi: 10.1038/nsmb.3135. Nat Struct Mol Biol. 2015. PMID: 26643844 No abstract available.
-
Autophagy proteins play cytoprotective and cytocidal roles in leucine starvation-induced cell death in Saccharomyces cerevisiae.Autophagy. 2012 May 1;8(5):731-8. doi: 10.4161/auto.19314. Epub 2012 Feb 24. Autophagy. 2012. PMID: 22361650 Free PMC article.
References
-
- Bates G. Huntingtin aggregation and toxicity in Huntington’s disease. Lancet. 2003;361:1642–1644. - PubMed
-
- Orr HT. Beyond the Qs in the polyglutamine diseases. Genes Dev. 2001;15:925–932. - PubMed
-
- Wanker EE. Protein aggregation and pathogenesis of Huntington’s disease: mechanisms and correlations. Biol Chem. 2000;381:937–942. - PubMed
-
- Gusella JF, MacDonald ME. Molecular genetics: unmasking polyglutamine triggers in neurodegenerative disease. Nat Rev Neurosci. 2000;1:109–115. - PubMed
-
- Albrecht M, Golatta M, Wullner U, Lengauer T. Structural and functional analysis of ataxin-2 and ataxin-3. Eur J Biochem. 2004;271:3155–3170. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

