Receiver domains control the active-state stoichiometry of Aquifex aeolicus sigma54 activator NtrC4, as revealed by electrospray ionization mass spectrometry
- PMID: 19699748
- PMCID: PMC2763505
- DOI: 10.1016/j.jmb.2009.08.033
Receiver domains control the active-state stoichiometry of Aquifex aeolicus sigma54 activator NtrC4, as revealed by electrospray ionization mass spectrometry
Abstract
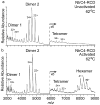
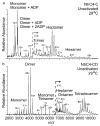
A common challenge with studies of proteins in vitro is determining which constructs and conditions are most physiologically relevant. sigma(54) activators are proteins that undergo regulated assembly to form an active ATPase ring that enables transcription by sigma(54)-polymerase. Previous studies of AAA(+) ATPase domains from sigma(54) activators have shown that some are heptamers, while others are hexamers. Because active oligomers assemble from off-state dimers, it was thought that even-numbered oligomers should dominate, and that heptamer formation would occur when individual domains of the activators, rather than the intact proteins, were studied. Here we present results from electrospray ionization mass spectrometry experiments characterizing the assembly states of intact NtrC4 (a sigma(54) activator from Aquifex aeolicus, an extreme thermophile), as well as its ATPase domain alone, and regulatory-ATPase and ATPase-DNA binding domain combinations. We show that the full-length and activated regulatory-ATPase proteins form hexamers, whereas the isolated ATPase domain, unactivated regulatory-ATPase, and ATPase-DNA binding domain form heptamers. Activation of the N-terminal regulatory domain is the key factor stabilizing the hexamer form of the ATPase, relative to the heptamer.
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

