Tetracycline-associated transcriptional regulation of transfer genes of the Bacteroides conjugative transposon CTnDOT
- PMID: 19700528
- PMCID: PMC2753023
- DOI: 10.1128/JB.00739-09
Tetracycline-associated transcriptional regulation of transfer genes of the Bacteroides conjugative transposon CTnDOT
Abstract
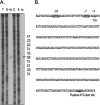
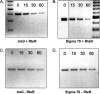
Many human colonic Bacteroides spp. harbor a conjugative transposon, CTnDOT, which carries two antibiotic resistance genes, tetQ and ermF. A distinctive feature of CTnDOT is that its excision and transfer are stimulated by tetracycline. Regulation of the genes responsible for excision has been described previously. We provide here the first characterization of the regulation of CTnDOT transfer (tra) genes. Reverse transcription-PCR analysis of the region containing the tra genes showed that these genes are regulated at the transcriptional level. Surprisingly, increased production of tra gene mRNA in tetracycline-stimulated cells was mediated by the proteins encoded by the excision genes. Previous studies have shown that expression of the excision gene operon is controlled by the regulatory protein RteC. Accordingly, it was possible that RteC was also regulating tra gene expression and that the excision proteins were only accessory proteins. However, placing the excision gene operon under the control of a heterologous promoter showed that the excision proteins alone could activate tra gene expression and that RteC was not directly involved. We also found a second level of tra gene control. The transfer of CTnDOT was inhibited by a DNA segment that included only a portion of the 3' end of one of the excision genes (exc). This segment contained a small open reading frame, rteR. By replacing the codons encoding the first two amino acids of the putative protein product of this open reading frame with stop codons, we showed that the rteR gene might encode a small regulatory RNA. RteR acted in trans to reduce the number of tra transcripts in a way that was independent of the excision proteins. The repressive effect of RteR was not the result of decreased stability of the tra mRNA. Instead, RteR appears to be modulating the level of tra gene expression in some more direct fashion. The complex regulatory system that controls and links the expression of CTnDOT excision and transfer genes may be designed to ensure stable maintenance of CTnDOT in nature by reducing the fitness toll it takes on the cell that harbors it.
Figures




Similar articles
-
The small RNA RteR inhibits transfer of the Bacteroides conjugative transposon CTnDOT.J Bacteriol. 2012 Oct;194(19):5228-36. doi: 10.1128/JB.00941-12. Epub 2012 Jul 20. J Bacteriol. 2012. PMID: 22821972 Free PMC article.
-
Regulation of excision genes of the Bacteroides conjugative transposon CTnDOT.J Bacteriol. 2005 Aug;187(16):5732-41. doi: 10.1128/JB.187.16.5732-5741.2005. J Bacteriol. 2005. PMID: 16077120 Free PMC article.
-
Tetracycline-related transcriptional regulation of the CTnDOT mobilization region.J Bacteriol. 2013 Dec;195(24):5431-8. doi: 10.1128/JB.00691-13. Epub 2013 Sep 27. J Bacteriol. 2013. PMID: 24078614 Free PMC article.
-
Regulation of CTnDOT conjugative transfer is a complex and highly coordinated series of events.mBio. 2013 Oct 29;4(6):e00569-13. doi: 10.1128/mBio.00569-13. mBio. 2013. PMID: 24169574 Free PMC article. Review.
-
The Integration and Excision of CTnDOT.Microbiol Spectr. 2015 Apr;3(2):MDNA3-0020-2014. doi: 10.1128/microbiolspec.MDNA3-0020-2014. Microbiol Spectr. 2015. PMID: 26104696 Free PMC article. Review.
Cited by
-
The Xis2d protein of CTnDOT binds to the intergenic region between the mob and tra operons.Plasmid. 2015 Sep;81:63-71. doi: 10.1016/j.plasmid.2015.07.002. Epub 2015 Jul 23. Plasmid. 2015. PMID: 26212728 Free PMC article.
-
Heterologous Complementation Studies With the YscX and YscY Protein Families Reveals a Specificity for Yersinia pseudotuberculosis Type III Secretion.Front Cell Infect Microbiol. 2018 Mar 16;8:80. doi: 10.3389/fcimb.2018.00080. eCollection 2018. Front Cell Infect Microbiol. 2018. PMID: 29616194 Free PMC article.
-
A high-resolution transcriptome map identifies small RNA regulation of metabolism in the gut microbe Bacteroides thetaiotaomicron.Nat Commun. 2020 Jul 16;11(1):3557. doi: 10.1038/s41467-020-17348-5. Nat Commun. 2020. PMID: 32678091 Free PMC article.
-
Integrative and conjugative elements: mosaic mobile genetic elements enabling dynamic lateral gene flow.Nat Rev Microbiol. 2010 Aug;8(8):552-63. doi: 10.1038/nrmicro2382. Epub 2010 Jul 5. Nat Rev Microbiol. 2010. PMID: 20601965 Review.
-
The Emerging Fish Pathogen Flavobacterium spartansii Isolated from Chinook Salmon: Comparative Genome Analysis and Molecular Manipulation.Front Microbiol. 2017 Nov 30;8:2339. doi: 10.3389/fmicb.2017.02339. eCollection 2017. Front Microbiol. 2017. PMID: 29250046 Free PMC article.
References
-
- Ball, T. B., F. A. Plummer, and K. T. Hayglass. 2003. Improved mRNA quantitation in LightCycler RT-PCR. Int. Arch. Allergy Immunol. 130:82-86. - PubMed
-
- Bayley, D. P., E. R. Rocha, and C. J. Smith. 2000. Analysis of cepA and other Bacteroides fragilis genes reveals a unique promoter structure. FEMS Microbiol. Lett. 193:149-154. - PubMed
-
- Bonheyo, G. T., B. D. Hund, N. B. Shoemaker, and A. A. Salyers. 2001. Transfer region of a Bacteroides conjugative transposon contains regulatory as well as structural genes. Plasmid 46:202-209. - PubMed
-
- Cheng, Q., Y. Sutanto, N. B. Shoemaker, J. F. Gardner, and A. A. Salyers. 2001. Identification of genes required for the excision of CTnDOT, a Bacteroides conjugative transposon. Mol. Microbiol. 41:625-632. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

