The coexistence of the nucleosome positioning code with the genetic code on eukaryotic genomes
- PMID: 19700771
- PMCID: PMC2770662
- DOI: 10.1093/nar/gkp689
The coexistence of the nucleosome positioning code with the genetic code on eukaryotic genomes
Abstract
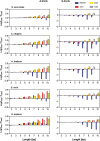
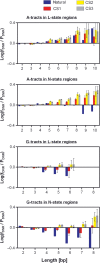
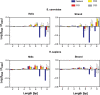
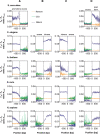
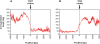
It is known that there are several codes residing simultaneously on the DNA double helix. The two best-characterized codes are the genetic code--the code for protein production, and the code for DNA packaging into nucleosomes. Since these codes have to coexist simultaneously on the same DNA region, both must be degenerate to allow this coexistence. A-tracts are homopolymeric stretches of several adjacent deoxyadenosines on one strand of the double helix, having unusual structural properties, which were shown to exclude nucleosomes and as such are instrumental in setting the translational positioning of DNA within nucleosomes. We observe, cross-kingdoms, a strong codon bias toward the avoidance of long A-tracts in exon regions, which enables the formation of high density of nucleosomes in these regions. Moreover, long A-tract avoidance is restricted exclusively to nucleosome-occupied exon regions. We show that this bias in codon usage is sufficient for enabling DNA organization within nucleosomes without constraints on the actual code for proteins. Thus, there is inter-dependency of the two major codes within DNA to allow their coexistence. Furthermore, we show that modulation of A-tract occurrences in exon versus non-exon regions may result in a unique alternation of the diameter of the '30-nm' fiber model.
Figures

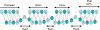
Similar articles
-
Poly(dA:dT) tracts: major determinants of nucleosome organization.Curr Opin Struct Biol. 2009 Feb;19(1):65-71. doi: 10.1016/j.sbi.2009.01.004. Epub 2009 Feb 7. Curr Opin Struct Biol. 2009. PMID: 19208466 Free PMC article. Review.
-
Long-range correlations between DNA bending sites: relation to the structure and dynamics of nucleosomes.J Mol Biol. 2002 Mar 1;316(4):903-18. doi: 10.1006/jmbi.2001.5363. J Mol Biol. 2002. PMID: 11884131
-
Modulation of cyclobutane pyrimidine dimer formation in a positioned nucleosome containing poly(dA.dT) tracts.Biochemistry. 1996 Jun 18;35(24):7705-14. doi: 10.1021/bi953011r. Biochemistry. 1996. PMID: 8672471
-
Re-cracking the nucleosome positioning code.Stat Appl Genet Mol Biol. 2008;7(1):Article14. doi: 10.2202/1544-6115.1367. Epub 2008 Apr 21. Stat Appl Genet Mol Biol. 2008. PMID: 18454729
-
DNA shape, genetic codes, and evolution.Curr Opin Struct Biol. 2011 Jun;21(3):342-7. doi: 10.1016/j.sbi.2011.03.002. Epub 2011 Mar 23. Curr Opin Struct Biol. 2011. PMID: 21439813 Free PMC article. Review.
Cited by
-
Adenine Enrichment at the Fourth CDS Residue in Bacterial Genes Is Consistent with Error Proofing for +1 Frameshifts.Mol Biol Evol. 2017 Dec 1;34(12):3064-3080. doi: 10.1093/molbev/msx223. Mol Biol Evol. 2017. PMID: 28961919 Free PMC article.
-
Regulation of chromatin folding by conformational variations of nucleosome linker DNA.Nucleic Acids Res. 2017 Sep 19;45(16):9372-9387. doi: 10.1093/nar/gkx562. Nucleic Acids Res. 2017. PMID: 28934465 Free PMC article.
-
Abundant oligonucleotides common to most bacteria.PLoS One. 2010 Mar 23;5(3):e9841. doi: 10.1371/journal.pone.0009841. PLoS One. 2010. PMID: 20352124 Free PMC article.
-
Proximal-end bias from in-vitro reconstituted nucleosomes and the result on downstream data analysis.PLoS One. 2021 Oct 21;16(10):e0258737. doi: 10.1371/journal.pone.0258737. eCollection 2021. PLoS One. 2021. PMID: 34673804 Free PMC article.
-
Predicting nucleosome binding motif set and analyzing their distributions around functional sites of human genes.Chromosome Res. 2012 Aug;20(6):685-98. doi: 10.1007/s10577-012-9305-0. Epub 2012 Jul 31. Chromosome Res. 2012. PMID: 22847645
References
-
- Trifonov EN. The multiple codes of nucleotide sequences. Bull. Math. Biol. 1989;51:417–432. - PubMed
-
- Nirenberg M, Caskey T, Marshall R, Brimacombe R, Kellogg D, Doctor B, Hatfield D, Levin J, Rottman F, Pestka S, et al. Cold Spring Harb. Symp. Quant. Biol. Vol. 31. Cold Spring Harbor, N.Y.: Cold Spring Harbor Laboratory; 1966. The RNA code and protein synthesis; pp. 11–24. - PubMed
-
- Rando OJ, Ahmad K. Rules and regulation in the primary structure of chromatin. Curr. Opin. Cell Biol. 2007;19:250–256. - PubMed
-
- Schnitzler GR. Control of nucleosome positions by DNA sequence and remodeling machines. Cell Biochem. Biophys. 2008;51:67–80. - PubMed
-
- Crick FH. Codon—anticodon pairing: the wobble hypothesis. J. Mol. Biol. 1966;19:548–555. - PubMed

