Mrd1p is required for release of base-paired U3 snoRNA within the preribosomal complex
- PMID: 19704003
- PMCID: PMC2772733
- DOI: 10.1128/MCB.00428-09
Mrd1p is required for release of base-paired U3 snoRNA within the preribosomal complex
Abstract
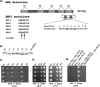
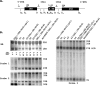
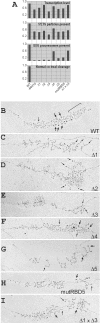
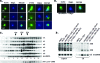
In eukaryotes, ribosomes are made from precursor rRNA (pre-rRNA) and ribosomal proteins in a maturation process that requires a large number of snoRNPs and processing factors. A fundamental problem is how the coordinated and productive folding of the pre-rRNA and assembly of successive pre-rRNA-protein complexes is achieved cotranscriptionally. The conserved protein Mrd1p, which contains five RNA binding domains (RBDs), is essential for processing events leading to small ribosomal subunit synthesis. We show that full function of Mrd1p requires all five RBDs and that the RBDs are functionally distinct and needed during different steps in processing. Mrd1p mutations trap U3 snoRNA in pre-rRNP complexes both in base-paired and non-base-paired interactions. A single essential RBD, RBD5, is involved in both types of interactions, but its conserved RNP1 motif is not needed for releasing the base-paired interactions. RBD5 is also required for the late pre-rRNP compaction preceding A(2) cleavage. Our results suggest that Mrd1p modulates successive conformational rearrangements within the pre-rRNP that influence snoRNA-pre-rRNA contacts and couple U3 snoRNA-pre-rRNA remodeling and late steps in pre-rRNP compaction that are essential for cleavage at A(0) to A(2). Mrd1p therefore coordinates key events in biosynthesis of small ribosome subunits.
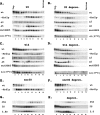
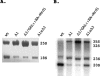
Figures
Similar articles
-
Linker 2 of the eukaryotic pre-ribosomal processing factor Mrd1p is an essential interdomain functionally coupled to upstream RNA Binding Domain 2 (RBD2).PLoS One. 2017 Apr 7;12(4):e0175506. doi: 10.1371/journal.pone.0175506. eCollection 2017. PLoS One. 2017. PMID: 28388671 Free PMC article.
-
Mrd1p binds to pre-rRNA early during transcription independent of U3 snoRNA and is required for compaction of the pre-rRNA into small subunit processomes.Nucleic Acids Res. 2008 Aug;36(13):4364-80. doi: 10.1093/nar/gkn384. Epub 2008 Jun 27. Nucleic Acids Res. 2008. PMID: 18586827 Free PMC article.
-
Mrd1p is required for processing of pre-rRNA and for maintenance of steady-state levels of 40 S ribosomal subunits in yeast.J Biol Chem. 2002 May 24;277(21):18431-9. doi: 10.1074/jbc.M112395200. Epub 2002 Mar 7. J Biol Chem. 2002. PMID: 11884397
-
Sno-capped: 5' ends of preribosomal RNAs are decorated with a U3 SnoRNP.Chem Biol. 2002 Jul;9(7):777-9. doi: 10.1016/s1074-5521(02)00171-0. Chem Biol. 2002. PMID: 12144919 Review.
-
Mpp10p, a new protein component of the U3 snoRNP required for processing of 18S rRNA precursors.Nucleic Acids Symp Ser. 1997;(36):64-7. Nucleic Acids Symp Ser. 1997. PMID: 9478208 Review.
Cited by
-
Multiple RNA interactions position Mrd1 at the site of the small subunit pseudoknot within the 90S pre-ribosome.Nucleic Acids Res. 2013 Jan;41(2):1178-90. doi: 10.1093/nar/gks1129. Epub 2012 Nov 27. Nucleic Acids Res. 2013. PMID: 23193268 Free PMC article.
-
Evolutionary conservation of the ribosomal biogenesis factor Rbm19/Mrd1: implications for function.PLoS One. 2012;7(9):e43786. doi: 10.1371/journal.pone.0043786. Epub 2012 Sep 12. PLoS One. 2012. PMID: 22984444 Free PMC article.
-
Linker 2 of the eukaryotic pre-ribosomal processing factor Mrd1p is an essential interdomain functionally coupled to upstream RNA Binding Domain 2 (RBD2).PLoS One. 2017 Apr 7;12(4):e0175506. doi: 10.1371/journal.pone.0175506. eCollection 2017. PLoS One. 2017. PMID: 28388671 Free PMC article.
-
Maturation of the 90S pre-ribosome requires Mrd1 dependent U3 snoRNA and 35S pre-rRNA structural rearrangements.Nucleic Acids Res. 2018 Apr 20;46(7):3692-3706. doi: 10.1093/nar/gky036. Nucleic Acids Res. 2018. PMID: 29373706 Free PMC article.
-
High-throughput RNA structure probing reveals critical folding events during early 60S ribosome assembly in yeast.Nat Commun. 2017 Sep 28;8(1):714. doi: 10.1038/s41467-017-00761-8. Nat Commun. 2017. PMID: 28959008 Free PMC article.
References
-
- Allain, F. H., D. E. Gilbert, P. Bouvet, and J. Feigon. 2000. Solution structure of the two N-terminal RNA-binding domains of nucleolin and NMR study of the interaction with its RNA target. J. Mol. Biol. 303:227-241. - PubMed
-
- Bleichert, F., and S. J. Baserga. 2007. The long unwinding road of RNA helicases. Mol. Cell 27:339-352. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
