The fission yeast homeodomain protein Yox1p binds to MBF and confines MBF-dependent cell-cycle transcription to G1-S via negative feedback
- PMID: 19714215
- PMCID: PMC2726434
- DOI: 10.1371/journal.pgen.1000626
The fission yeast homeodomain protein Yox1p binds to MBF and confines MBF-dependent cell-cycle transcription to G1-S via negative feedback
Abstract
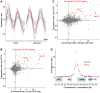
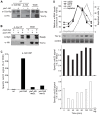
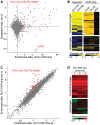
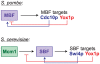
The regulation of the G1- to S-phase transition is critical for cell-cycle progression. This transition is driven by a transient transcriptional wave regulated by transcription factor complexes termed MBF/SBF in yeast and E2F-DP in mammals. Here we apply genomic, genetic, and biochemical approaches to show that the Yox1p homeodomain protein of fission yeast plays a critical role in confining MBF-dependent transcription to the G1/S transition of the cell cycle. The yox1 gene is an MBF target, and Yox1p accumulates and preferentially binds to MBF-regulated promoters, via the MBF components Res2p and Nrm1p, when they are transcriptionally repressed during the cell cycle. Deletion of yox1 results in constitutively high transcription of MBF target genes and loss of their cell cycle-regulated expression, similar to deletion of nrm1. Genome-wide location analyses of Yox1p and the MBF component Cdc10p reveal dozens of genes whose promoters are bound by both factors, including their own genes and histone genes. In addition, Cdc10p shows promiscuous binding to other sites, most notably close to replication origins. This study establishes Yox1p as a new regulatory MBF component in fission yeast, which is transcriptionally induced by MBF and in turn inhibits MBF-dependent transcription. Yox1p may function together with Nrm1p to confine MBF-dependent transcription to the G1/S transition of the cell cycle via negative feedback. Compared to the orthologous budding yeast Yox1p, which indirectly functions in a negative feedback loop for cell-cycle transcription, similarities but also notable differences in the wiring of the regulatory circuits are evident.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
Phosphorylation of the MBF repressor Yox1p by the DNA replication checkpoint keeps the G1/S cell-cycle transcriptional program active.PLoS One. 2011 Feb 16;6(2):e17211. doi: 10.1371/journal.pone.0017211. PLoS One. 2011. PMID: 21359180 Free PMC article.
-
Gcn5-mediated acetylation at MBF-regulated promoters induces the G1/S transcriptional wave.Nucleic Acids Res. 2019 Sep 19;47(16):8439-8451. doi: 10.1093/nar/gkz561. Nucleic Acids Res. 2019. PMID: 31260531 Free PMC article.
-
SWI/SNF and RSC remodeler complexes bind to MBF-dependent genes.Cell Cycle. 2021 Dec;20(24):2652-2661. doi: 10.1080/15384101.2021.2008203. Epub 2021 Nov 29. Cell Cycle. 2021. PMID: 34843421 Free PMC article.
-
Down-regulation of Cdk1 activity in G1 coordinates the G1/S gene expression programme with genome replication.Curr Genet. 2019 Jun;65(3):685-690. doi: 10.1007/s00294-018-00926-y. Epub 2019 Jan 24. Curr Genet. 2019. PMID: 30680437 Review.
-
Isolation and characterization of fission yeast genes involved in transcription regulation of cell cycle events (a short communication).Acta Microbiol Immunol Hung. 2002;49(2-3):285-7. doi: 10.1556/AMicr.49.2002.2-3.16. Acta Microbiol Immunol Hung. 2002. PMID: 12109160 Review. No abstract available.
Cited by
-
Comprehensive proteomics analysis reveals new substrates and regulators of the fission yeast clp1/cdc14 phosphatase.Mol Cell Proteomics. 2013 May;12(5):1074-86. doi: 10.1074/mcp.M112.025924. Epub 2013 Jan 7. Mol Cell Proteomics. 2013. PMID: 23297348 Free PMC article.
-
Genetic-interaction screens uncover novel biological roles and regulators of transcription factors in fission yeast.G3 (Bethesda). 2022 Aug 25;12(9):jkac194. doi: 10.1093/g3journal/jkac194. G3 (Bethesda). 2022. PMID: 35924983 Free PMC article.
-
The DNA damage and the DNA replication checkpoints converge at the MBF transcription factor.Mol Biol Cell. 2013 Nov;24(21):3350-7. doi: 10.1091/mbc.E13-05-0257. Epub 2013 Sep 4. Mol Biol Cell. 2013. PMID: 24006488 Free PMC article.
-
Consensus clustering for Bayesian mixture models.BMC Bioinformatics. 2022 Jul 21;23(1):290. doi: 10.1186/s12859-022-04830-8. BMC Bioinformatics. 2022. PMID: 35864476 Free PMC article.
-
Nrm1 is a bistable switch connecting cell cycle progression to transcriptional control.EMBO Rep. 2025 Aug 29. doi: 10.1038/s44319-025-00566-7. Online ahead of print. EMBO Rep. 2025. PMID: 40883509
References
-
- Bähler J. Cell-cycle control of gene expression in budding and fission yeast. Annu Rev Genet. 2005;39:69–94. - PubMed
-
- Breeden LL. Periodic transcription: a cycle within a cycle. Curr Biol. 2003;13:R31–R38. - PubMed
-
- Wittenberg C, Reed SI. Cell cycle-dependent transcription in yeast: promoters, transcription factors, and transcriptomes. Oncogene. 2005;24:2746–2755. - PubMed
-
- Nevins JR. The Rb/E2F pathway and cancer. Hum Mol Genet. 2001;10:699–703. - PubMed
-
- Sherr CJ, McCormick F. The RB and p53 pathways in cancer. Cancer Cell. 2002;2:103–112. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

