Copy number variation in human health, disease, and evolution
- PMID: 19715442
- PMCID: PMC4472309
- DOI: 10.1146/annurev.genom.9.081307.164217
Copy number variation in human health, disease, and evolution
Abstract
Copy number variation (CNV) is a source of genetic diversity in humans. Numerous CNVs are being identified with various genome analysis platforms, including array comparative genomic hybridization (aCGH), single nucleotide polymorphism (SNP) genotyping platforms, and next-generation sequencing. CNV formation occurs by both recombination-based and replication-based mechanisms and de novo locus-specific mutation rates appear much higher for CNVs than for SNPs. By various molecular mechanisms, including gene dosage, gene disruption, gene fusion, position effects, etc., CNVs can cause Mendelian or sporadic traits, or be associated with complex diseases. However, CNV can also represent benign polymorphic variants. CNVs, especially gene duplication and exon shuffling, can be a predominant mechanism driving gene and genome evolution.
Figures
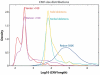
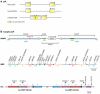


References
-
- Aitman TJ, Dong R, Vyse TJ, Norsworthy PJ, Johnson MD, et al. Copy number polymorphism in Fcgr3 predisposes to glomerulonephritis in rats and humans. Nature. 2006;439:851–55. - PubMed
-
- Babushok DV, Kazazian HH., Jr. Progress in understanding the biology of the human mutagen LINE-1. Hum. Mutat. 2007;28:527–39. - PubMed
-
- Bailey A, Phillips W, Rutter M. Autism: towards an integration of clinical, genetic, neuropsychological, and neurobiological perspectives. J. Child Psychol. Psychiatry. 1996;37:89–126. - PubMed
-
- Bailey JA, Gu Z, Clark RA, Reinert K, Samonte RV, et al. Recent segmental duplications in the human genome. Science. 2002;297:1003–7. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

