Comparative genomics reveals mechanism for short-term and long-term clonal transitions in pandemic Vibrio cholerae
- PMID: 19720995
- PMCID: PMC2741270
- DOI: 10.1073/pnas.0907787106
Comparative genomics reveals mechanism for short-term and long-term clonal transitions in pandemic Vibrio cholerae
Abstract
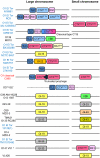
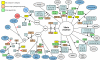
Vibrio cholerae, the causative agent of cholera, is a bacterium autochthonous to the aquatic environment, and a serious public health threat. V. cholerae serogroup O1 is responsible for the previous two cholera pandemics, in which classical and El Tor biotypes were dominant in the sixth and the current seventh pandemics, respectively. Cholera researchers continually face newly emerging and reemerging pathogenic clones carrying diverse combinations of phenotypic and genotypic properties, which significantly hampered control of the disease. To elucidate evolutionary mechanisms governing genetic diversity of pandemic V. cholerae, we compared the genome sequences of 23 V. cholerae strains isolated from a variety of sources over the past 98 years. The genome-based phylogeny revealed 12 distinct V. cholerae lineages, of which one comprises both O1 classical and El Tor biotypes. All seventh pandemic clones share nearly identical gene content. Using analogy to influenza virology, we define the transition from sixth to seventh pandemic strains as a "shift" between pathogenic clones belonging to the same O1 serogroup, but from significantly different phyletic lineages. In contrast, transition among clones during the present pandemic period is characterized as a "drift" between clones, differentiated mainly by varying composition of laterally transferred genomic islands, resulting in emergence of variants, exemplified by V. cholerae O139 and V. cholerae O1 El Tor hybrid clones. Based on the comparative genomics it is concluded that V. cholerae undergoes extensive genetic recombination via lateral gene transfer, and, therefore, genome assortment, not serogroup, should be used to define pathogenic V. cholerae clones.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

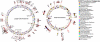
References
-
- Chatterjee SN, Chaudhuri K. Lipopolysaccharides of Vibrio cholerae. I. Physical and chemical characterization. Biochim Biophys Acta. 2003;1639:65–79. - PubMed
-
- Colwell RR. Global climate and infectious disease: The cholera paradigm. Science. 1996;274:2025–2031. - PubMed
-
- Ramamurthy T, et al. Emergence of novel strain of Vibrio cholerae with epidemic potential in southern and eastern India. Lancet. 1993;341:703–704. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

