Single nucleotide polymorphisms on the road to strain differentiation in Mycobacterium ulcerans
- PMID: 19726608
- PMCID: PMC2772594
- DOI: 10.1128/JCM.00761-09
Single nucleotide polymorphisms on the road to strain differentiation in Mycobacterium ulcerans
Abstract
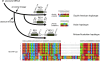
The genomic fine-typing of strains of Mycobacterium ulcerans, the causative agent of the emerging human disease Buruli ulcer, is difficult due to the clonal population structure of geographical lineages. Although large sequence polymorphisms (LSPs) resulted in the clustering of patient isolates originating from across the globe, differentiation of strains within continents using conventional typing methods is very limited. In this study, we analyzed M. ulcerans LSP haplotype-specific insertion sequence elements among 83 M. ulcerans strains and identified single nucleotide polymorphisms (SNPs) that differentiate between regional strains. This is the first genetic discrimination based on SNPs of M. ulcerans strains from African countries where Buruli ulcer is endemic, resulting in the highest geographic resolution of genotyping so far. The findings support the concept of genome-wide SNP analyses as tools to study the epidemiology and evolution of M. ulcerans at a local level.
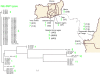
Figures
Similar articles
-
Lack of insertional-deletional polymorphism in a collection of Mycobacterium ulcerans isolates from Ghanaian Buruli ulcer patients.J Clin Microbiol. 2009 Nov;47(11):3640-6. doi: 10.1128/JCM.00760-09. Epub 2009 Sep 2. J Clin Microbiol. 2009. PMID: 19726605 Free PMC article.
-
Insertion sequence element single nucleotide polymorphism typing provides insights into the population structure and evolution of Mycobacterium ulcerans across Africa.Appl Environ Microbiol. 2014 Feb;80(3):1197-209. doi: 10.1128/AEM.02774-13. Epub 2013 Dec 2. Appl Environ Microbiol. 2014. PMID: 24296504 Free PMC article.
-
Single nucleotide polymorphism typing of Mycobacterium ulcerans reveals focal transmission of buruli ulcer in a highly endemic region of Ghana.PLoS Negl Trop Dis. 2010 Jul 20;4(7):e751. doi: 10.1371/journal.pntd.0000751. PLoS Negl Trop Dis. 2010. PMID: 20652033 Free PMC article.
-
The genome, evolution and diversity of Mycobacterium ulcerans.Infect Genet Evol. 2012 Apr;12(3):522-9. doi: 10.1016/j.meegid.2012.01.018. Epub 2012 Jan 28. Infect Genet Evol. 2012. PMID: 22306192 Review.
-
Buruli ulcer: reductive evolution enhances pathogenicity of Mycobacterium ulcerans.Nat Rev Microbiol. 2009 Jan;7(1):50-60. doi: 10.1038/nrmicro2077. Nat Rev Microbiol. 2009. PMID: 19079352 Review.
Cited by
-
Lack of insertional-deletional polymorphism in a collection of Mycobacterium ulcerans isolates from Ghanaian Buruli ulcer patients.J Clin Microbiol. 2009 Nov;47(11):3640-6. doi: 10.1128/JCM.00760-09. Epub 2009 Sep 2. J Clin Microbiol. 2009. PMID: 19726605 Free PMC article.
-
Sero-epidemiology as a tool to screen populations for exposure to Mycobacterium ulcerans.PLoS Negl Trop Dis. 2012 Jan;6(1):e1460. doi: 10.1371/journal.pntd.0001460. Epub 2012 Jan 10. PLoS Negl Trop Dis. 2012. PMID: 22253937 Free PMC article.
-
Insertion sequence element single nucleotide polymorphism typing provides insights into the population structure and evolution of Mycobacterium ulcerans across Africa.Appl Environ Microbiol. 2014 Feb;80(3):1197-209. doi: 10.1128/AEM.02774-13. Epub 2013 Dec 2. Appl Environ Microbiol. 2014. PMID: 24296504 Free PMC article.
-
Genotyping Tools for Mycobacterium ulcerans-Drawbacks and Future Prospects.Mycobact Dis. 2014 May 5;4(2):1000149. doi: 10.4172/2161-1068.1000149. Mycobact Dis. 2014. PMID: 24900947 Free PMC article.
-
Single nucleotide polymorphism typing of Mycobacterium ulcerans reveals focal transmission of buruli ulcer in a highly endemic region of Ghana.PLoS Negl Trop Dis. 2010 Jul 20;4(7):e751. doi: 10.1371/journal.pntd.0000751. PLoS Negl Trop Dis. 2010. PMID: 20652033 Free PMC article.
References
-
- Ablordey, A., P. A. Fonteyne, P. Stragier, P. Vandamme, and F. Portaels. 2007. Identification of a new variable number tandem repeat locus in Mycobacterium ulcerans for potential strain discrimination among African isolates. Clin. Microbiol. Infect. 13:734-736. - PubMed
-
- Brosch, R., S. V. Gordon, M. Marmiesse, P. Brodin, C. Buchrieser, K. Eiglmeier, T. Garnier, C. Gutierrez, G. Hewinson, K. Kremer, L. M. Parsons, A. S. Pym, S. Samper, D. van Soolingen, and S. T. Cole. 2002. A new evolutionary scenario for the Mycobacterium tuberculosis complex. Proc. Natl. Acad. Sci. USA 99:3684-3689. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

