Aedes aegypti cadherin serves as a putative receptor of the Cry11Aa toxin from Bacillus thuringiensis subsp. israelensis
- PMID: 19732034
- PMCID: PMC3702384
- DOI: 10.1042/BJ20090730
Aedes aegypti cadherin serves as a putative receptor of the Cry11Aa toxin from Bacillus thuringiensis subsp. israelensis
Abstract
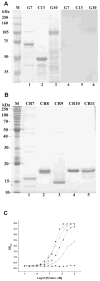
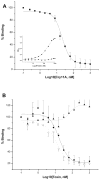
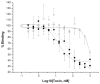
Cry11Aa of Bacillus thuringiensis subsp. israelensis is the most active toxin to Aedes aegypti in this strain. We previously reported that, in addition to a 65 kDa GPI (glycosylphosphatidylinositol)-anchored ALP (alkaline phosphatase), the toxin also binds a 250 kDa membrane protein. Since this protein is the same size as cadherin, which in lepidopteran insects is an important Cry toxin receptor, we developed an anti-AaeCad antibody. This antibody detects a 250 kDa protein in immunoblots of larval BBMVs (brush border membrane vesicles). The antibody inhibits Cry11Aa toxin binding to BBMVs and immunolocalizes the cadherin protein to apical membranes of distal and proximal caecae and posterior midgut epithelial cells. This localization is consistent with areas to which Cry11Aa toxin binds and causes pathogenicity. Therefore, the full-length Aedes cadherin cDNA was isolated from Aedes larvae and partial overlapping fragments that covered the entire protein were expressed in Escherichia coli. Using toxin overlay assays, we showed that one cadherin fragment, which contains CR7-11 (cadherin repeats 7-11), bound Cry11Aa and this binding was primarily through toxin domain II loops alpha8 and 2. Cadherin repeats CR8-11 but not CR7 bound Cry11Aa under non-denaturing conditions. Cry11Aa bound the cadherin fragment with high affinity with an apparent Kd of 16.7 nM. Finally we showed that this Cry11Aa-binding site could also be competed by Cry11Ba and Cry4Aa but not Cry4Ba. These results indicate that Aedes cadherin is possibly a receptor for Cry11A and, together with its ability to bind an ALP, suggest a similar mechanism of toxin action as previously proposed for lepidopteran insects.
Figures



Similar articles
-
Cadherin binding is not a limiting step for Bacillus thuringiensis subsp. israelensis Cry4Ba toxicity to Aedes aegypti larvae.Biochem J. 2012 May 1;443(3):711-7. doi: 10.1042/BJ20111579. Biochem J. 2012. PMID: 22329749 Free PMC article.
-
A 104 kDa Aedes aegypti aminopeptidase N is a putative receptor for the Cry11Aa toxin from Bacillus thuringiensis subsp. israelensis.Insect Biochem Mol Biol. 2013 Dec;43(12):1201-8. doi: 10.1016/j.ibmb.2013.09.007. Epub 2013 Oct 12. Insect Biochem Mol Biol. 2013. PMID: 24128608 Free PMC article.
-
Cadherin, alkaline phosphatase, and aminopeptidase N as receptors of Cry11Ba toxin from Bacillus thuringiensis subsp. jegathesan in Aedes aegypti.Appl Environ Microbiol. 2011 Jan;77(1):24-31. doi: 10.1128/AEM.01852-10. Epub 2010 Oct 29. Appl Environ Microbiol. 2011. PMID: 21037295 Free PMC article.
-
Bacillus thuringiensis subsp. israelensis and its dipteran-specific toxins.Toxins (Basel). 2014 Mar 28;6(4):1222-43. doi: 10.3390/toxins6041222. Toxins (Basel). 2014. PMID: 24686769 Free PMC article. Review.
-
Employing phage display to study the mode of action of Bacillus thuringiensis Cry toxins.Peptides. 2008 Feb;29(2):324-9. doi: 10.1016/j.peptides.2007.07.035. Epub 2007 Dec 14. Peptides. 2008. PMID: 18226423 Free PMC article. Review.
Cited by
-
Role of Lectin in the Response of Aedes aegypti Against Bt Toxin.Front Immunol. 2022 May 13;13:898198. doi: 10.3389/fimmu.2022.898198. eCollection 2022. Front Immunol. 2022. PMID: 35634312 Free PMC article. Review.
-
Bacillus thuringiensis: A story of a successful bioinsecticide.Insect Biochem Mol Biol. 2011 Jul;41(7):423-31. doi: 10.1016/j.ibmb.2011.02.006. Epub 2011 Mar 2. Insect Biochem Mol Biol. 2011. PMID: 21376122 Free PMC article. Review.
-
An Intramolecular Salt Bridge in Bacillus thuringiensis Cry4Ba Toxin Is Involved in the Stability of Helix α-3, Which Is Needed for Oligomerization and Insecticidal Activity.Appl Environ Microbiol. 2017 Sep 29;83(20):e01515-17. doi: 10.1128/AEM.01515-17. Print 2017 Oct 15. Appl Environ Microbiol. 2017. PMID: 28802270 Free PMC article.
-
Aedes cadherin mediates the in vivo toxicity of the Cry11Aa toxin to Aedes aegypti.Peptides. 2015 Jun;68:140-147. doi: 10.1016/j.peptides.2014.07.015. Epub 2014 Jul 23. Peptides. 2015. PMID: 25064814 Free PMC article.
-
In silico models for predicting vector control chemicals targeting Aedes aegypti.SAR QSAR Environ Res. 2014;25(10):805-35. doi: 10.1080/1062936X.2014.958291. Epub 2014 Oct 2. SAR QSAR Environ Res. 2014. PMID: 25275884 Free PMC article. Review.
References
-
- Kyle JL, Harris E. Global spread and persistence of dengue. Annu Rev Microbiol. 2008;62:71–92. - PubMed
-
- Bravo A, Gill SS, Soberon M. Bacillus thuringiensis: mechanisms and use. In: Gilbert LI, Kostas I, Gill SS, editors. Comprehensive Molecular Insect Science. Vol. 6. Elsevier; 2005. pp. 175–205.
-
- Dai SM, Gill SS. In vitro and in vivo proteolysis of the Bacillus thuringiensis subsp. israelensis CryIVD protein by Culex quinquefasciatus larval midgut proteases. Insect Biochem Mol Biol. 1993;23:273–283. - PubMed
-
- Yamagiwa M, Sakagawa K, Sakai H. Functional analysis of two processed fragments of Bacillus thuringiensis Cry11A toxin. Biosci Biotechnol Biochem. 2004;68:523–528. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

