Small RNAs and small proteins involved in resistance to cell envelope stress and acid shock in Escherichia coli: analysis of a bar-coded mutant collection
- PMID: 19734312
- PMCID: PMC2798238
- DOI: 10.1128/JB.00873-09
Small RNAs and small proteins involved in resistance to cell envelope stress and acid shock in Escherichia coli: analysis of a bar-coded mutant collection
Abstract
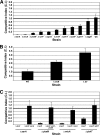
More than 80 small regulatory RNAs (sRNAs) and 60 proteins of 16 to 50 amino acids (small proteins) are encoded in the Escherichia coli genome. The vast majority of the corresponding genes have no known function. We screened 125 DNA bar-coded mutants to identify novel cell envelope stress and acute acid shock phenotypes associated with deletions of genes coding for sRNAs and small proteins. Nine deletion mutants (ssrA, micA, ybaM, ryeF, yqcG, sroH, ybhT, yobF, and glmY) were sensitive to cell envelope stress and two were resistant (rybB and blr). Deletion mutants of genes coding for four small proteins (yqgB, mgrB, yobF, and yceO) were sensitive to acute acid stress. We confirmed each of these phenotypes in one-on-one competition assays against otherwise-wild-type lacZ mutant cells. A more detailed investigation of the SsrA phenotype suggests that ribosome release is critical for resistance to cell envelope stress. The bar-coded deletion collection we generated can be screened for sensitivity or resistance to virtually any stress condition.
Figures
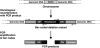



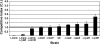

Comment in
-
Global approaches for finding small RNA and small open reading frame functions.J Bacteriol. 2010 Jan;192(1):26-8. doi: 10.1128/JB.01316-09. J Bacteriol. 2010. PMID: 19854892 Free PMC article. No abstract available.
References
-
- Altuvia, S. 2007. Identification of bacterial small non-coding RNAs: experimental approaches. Curr. Opin. Microbiol. 10:257-261. - PubMed
-
- Bos, M. P., V. Robert, and J. Tommassen. 2007. Biogenesis of the Gram-negative bacterial outer membrane. Annu. Rev. Microbiol. 61:191-214. - PubMed
-
- Boyle, S. M., G. D. Markham, E. W. Hafner, J. M. Wright, H. Tabor, and C. W. Tabor. 1984. Expression of the cloned genes encoding the putrescine biosynthetic enzymes and methionine adenosyltransferase of Escherichia coli (speA, speB, speC and metK). Gene 30:129-136. - PubMed
-
- Butland, G., M. Babu, J. J. Diaz-Mejia, F. Bohdana, S. Phanse, B. Gold, W. Yang, J. Li, A. G. Gagarinova, O. Pogoutse, H. Mori, B. L. Wanner, H. Lo, J. Wasniewski, C. Christopolous, M. Ali, P. Venn, A. Safavi-Naini, N. Sourour, S. Caron, J. Y. Choi, L. Laigle, A. Nazarians-Armavil, A. Deshpande, S. Joe, K. A. Datsenko, N. Yamamoto, B. J. Andrews, C. Boone, H. Ding, B. Sheikh, G. Moreno-Hagelseib, J. F. Greenblatt, and A. Emili. 2008. eSGA: E. coli synthetic genetic array analysis. Nat. Methods 5:789-795. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

