A robust two-dimensional separation for top-down tandem mass spectrometry of the low-mass proteome
- PMID: 19747844
- PMCID: PMC2830800
- DOI: 10.1016/j.jasms.2009.08.001
A robust two-dimensional separation for top-down tandem mass spectrometry of the low-mass proteome
Abstract
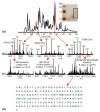
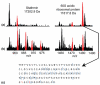
For fractionation of intact proteins by molecular weight (MW), a sharply improved two-dimensional (2D) separation is presented to drive reproducible and robust fractionation before top-down mass spectrometry of complex mixtures. The "GELFrEE" (i.e., gel-eluted liquid fraction entrapment electrophoresis) approach is implemented by use of Tris-glycine and Tris-tricine gel systems applied to human cytosolic and nuclear extracts from HeLa S3 cells, to achieve a MW-based fractionation of proteins from 5 to >100 kDa in 1 h. For top-down tandem mass spectroscopy (MS/MS) of the low-mass proteome (5-25 kDa), between 5 and 8 gel-elution (GE) fractions are sampled by nanocapillary-LC-MS/MS with 12 or 14.5 tesla Fourier transform ion cyclotron resonance (FT-ICR) mass spectrometers. Single injections give about 40 detectable proteins, about half of which yield automated ProSight identifications. Reproducibility metrics of the system are presented, along with comparative analysis of protein targets in mitotic versus asynchronous cells. We forward this basic 2D approach to facilitate wider implementation of top-down mass spectrometry and a variety of other protein separation and/or characterization approaches.
Figures


References
-
- Kelleher NL, Costello CA, Begley TP, McLafferty FW. Thiaminase-I (42-kDa) Heterogeneity, Sequence Refinement, and Active-Site Location from High-Resolution Tandem Mass-Spectrometry. J. Am. Soc. Mass Spectrom. 1995;6:981–984. - PubMed
-
- Kelleher NL, Lin HY, Valaskovic GA, Aaserud DJ, Fridriksson EK, McLafferty FW. Top Down versus Bottom Up Protein Characterization by Tandem High-Resolution Mass Spectrometry. J. Am. Chem. Soc. 1999;121:806–812.
-
- McLafferty FW. High-Resolution Tandem FT Mass-Spectrometry above 10-kDa. Acc. Chem. Res. 1994;27:379–386.
-
- Chong BE, Yan F, Lubman DM, Miller FR. Chromatofocusing Nonporous Reversed-Phase High-Performance Liquid Chromatography/Electrospray Ionization Time-of-Flight Mass Spectrometry of Proteins from Human Breast Cancer Whole Cell Lysates: A Novel Two-Dimensional Liquid Chromatography/Mass Spectrometry Method. Rapid Commun. Mass Spectrom. 2001;15:291–296. - PubMed
-
- Lubman DM, Kachman MT, Wang HX, Gong SY, Yan F, Hamler RL, O’Neil KA, Zhu K, Buchanan NS, Barder TJ. Two-Dimensional Liquid Separations—Mass Mapping of Proteins from Human Cancer Cell Lysates. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2002;782:183–196. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

