The nuclear pore complex has entered the atomic age
- PMID: 19748337
- PMCID: PMC2776643
- DOI: 10.1016/j.str.2009.07.014
The nuclear pore complex has entered the atomic age
Abstract
Nuclear pore complexes (NPCs) perforate the nuclear envelope and represent the exclusive passageway into and out of the nucleus of the eukaryotic cell. Apart from their essential transport function, components of the NPC have important, direct roles in nuclear organization and in gene regulation. Because of its central role in cell biology, it is of considerable interest to determine the NPC structure at atomic resolution. The complexity of these large, 40-60 MDa protein assemblies has for decades limited such structural studies. More recently, exploiting the intrinsic modularity of the NPC, structural biologists are making progress toward understanding this nanomachine in molecular detail. Structures of building blocks of the stable, architectural scaffold of the NPC have been solved, and distinct models for their assembly proposed. Here we review the status of the field and lay out the challenges and the next steps toward a full understanding of the NPC at atomic resolution.
Figures

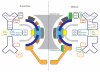




References
-
- Akhtar A, Gasser SM. The nuclear envelope and transcriptional control. Nat. Rev. Genet. 2007;8:507–517. - PubMed
-
- Alber F, Dokudovskaya S, Veenhoff LM, Zhang W, Kipper J, Devos D, Suprapto A, Karni-Schmidt O, Williams R, Chait BT, et al. Determining the architectures of macromolecular assemblies. Nature. 2007a;450:683–694. - PubMed
-
- Alber F, Dokudovskaya S, Veenhoff LM, Zhang W, Kipper J, Devos D, Suprapto A, Karni-Schmidt O, Williams R, Chait BT, et al. The molecular architecture of the nuclear pore complex. Nature. 2007b;450:695–701. - PubMed
-
- Alber F, Forster F, Korkin D, Topf M, Sali A. Integrating diverse data for structure determination of macromolecular assemblies. Annu. Rev. Biochem. 2008;77:443–477. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

