Structure of the Yeast DEAD box protein Mss116p reveals two wedges that crimp RNA
- PMID: 19748356
- PMCID: PMC2939728
- DOI: 10.1016/j.molcel.2009.07.032
Structure of the Yeast DEAD box protein Mss116p reveals two wedges that crimp RNA
Abstract
The yeast DEAD box protein Mss116p is a general RNA chaperone that functions in mitochondrial group I and II intron splicing, translational activation, and RNA end processing. Here we determined high-resolution X-ray crystal structures of Mss116p complexed with an RNA oligonucleotide and ATP analogs AMP-PNP, ADP-BeF(3)(-), or ADP-AlF(4)(-). The structures show the entire helicase core acting together with a functionally important C-terminal extension. In all structures, the helicase core is in a closed conformation with a wedge alpha helix bending RNA 3' of the central bound nucleotides, as in previous DEAD box protein structures. Notably, Mss116p's C-terminal extension also bends RNA 5' of the central nucleotides, resulting in RNA crimping. Despite reported functional differences, we observe few structural changes in ternary complexes with different ATP analogs. The structures constrain models of DEAD box protein function and reveal a strand separation mechanism in which a protein uses two wedges to act as a molecular crimper.
Figures




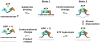
References
-
- Andersen CB, Ballut L, Johansen JS, Chamieh H, Nielsen KH, Oliveira CL, Pedersen JS, Séraphin B, Le Hir H, Andersen GR. Structure of the exon junction core complex with a trapped DEAD-box ATPase bound to RNA. Science. 2006;313:1968–1972. - PubMed
-
- Bono F, Ebert J, Lorentzen E, Conti E. The crystal structure of the exon junction complex reveals how it maintains a stable grip on mRNA. Cell. 2006;126:713–725. - PubMed
-
- CCP4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr D Biol Crystallogr. 1994;50:760–763. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

