Genomic imprinting: employing and avoiding epigenetic processes
- PMID: 19759261
- PMCID: PMC2751984
- DOI: 10.1101/gad.1841409
Genomic imprinting: employing and avoiding epigenetic processes
Abstract
Genomic imprinting refers to an epigenetic mark that distinguishes parental alleles and results in a monoallelic, parental-specific expression pattern in mammals. Few phenomena in nature depend more on epigenetic mechanisms while at the same time evading them. The alleles of imprinted genes are marked epigenetically at discrete elements termed imprinting control regions (ICRs) with their parental origin in gametes through the use of DNA methylation, at the very least. Imprinted gene expression is subsequently maintained using noncoding RNAs, histone modifications, insulators, and higher-order chromatin structure. Avoidance is manifest when imprinted genes evade the genome-wide reprogramming that occurs after fertilization and remain marked with their parental origin. This review summarizes what is known about the establishment and maintenance of imprinting marks and discusses the mechanisms of imprinting in clusters. Additionally, the evolution of imprinted gene clusters is described. While considerable information regarding epigenetic control of imprinting has been obtained recently, much remains to be learned.
Figures
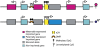
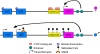

References
-
- Babak T, Deveale B, Armour C, Raymond C, Cleary MA, van der Kooy D, Johnson JM, Lim LP. Global survey of genomic imprinting by transcriptome sequencing. Curr Biol. 2008;18:1735–1741. - PubMed
-
- Barski A, Cuddapah S, Cui K, Roh TY, Schones DE, Wang Z, Wei G, Chepelev I, Zhao K. High-resolution profiling of histone methylations in the human genome. Cell. 2007;129:823–837. - PubMed
-
- Bartolomei MS, Zemel S, Tilghman SM. Parental imprinting of the mouse H19 gene. Nature. 1991;351:153–155. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
