Contextual extracellular cues promote tumor cell EMT and metastasis by regulating miR-200 family expression
- PMID: 19759262
- PMCID: PMC2751985
- DOI: 10.1101/gad.1820209
Contextual extracellular cues promote tumor cell EMT and metastasis by regulating miR-200 family expression
Abstract
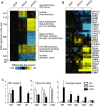
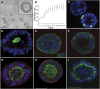
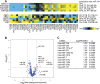
Metastatic disease is a primary cause of cancer-related death, and factors governing tumor cell metastasis have not been fully elucidated. Here, we address this question by using tumor cell lines derived from mice that develop metastatic lung adenocarcinoma owing to expression of mutant K-ras and p53. Despite having widespread somatic genetic alterations, the metastasis-prone tumor cells retained a marked plasticity. They transited reversibly between epithelial and mesenchymal states, forming highly polarized epithelial spheres in three-dimensional culture that underwent epithelial-to-mesenchymal transition (EMT) following treatment with transforming growth factor-beta or injection into syngeneic mice. This transition was entirely dependent on the microRNA (miR)-200 family, which decreased during EMT. Forced expression of miR-200 abrogated the capacity of these tumor cells to undergo EMT, invade, and metastasize, and conferred transcriptional features of metastasis-incompetent tumor cells. We conclude that tumor cell metastasis is regulated by miR-200 expression, which changes in response to contextual extracellular cues.
Figures




References
-
- Bartel DP. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297. - PubMed
-
- Bracken CP, Gregory PA, Kolesnikoff N, Bert AG, Wang J, Shannon MF, Goodall GJ. A double-negative feedback loop between ZEB1-SIP1 and the microRNA-200 family regulates epithelial-mesenchymal transition. Cancer Res. 2008;68:7846–7854. - PubMed
-
- Bussemaker HJ, Li H, Siggia ED. Regulatory element detection using correlation with expression. Nat Genet. 2001;27:167–171. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
