Genetic analysis of variation in human meiotic recombination
- PMID: 19763160
- PMCID: PMC2730532
- DOI: 10.1371/journal.pgen.1000648
Genetic analysis of variation in human meiotic recombination
Abstract
The number of recombination events per meiosis varies extensively among individuals. This recombination phenotype differs between female and male, and also among individuals of each gender. In this study, we used high-density SNP genotypes of over 2,300 individuals and their offspring in two datasets to characterize recombination landscape and to map the genetic variants that contribute to variation in recombination phenotypes. We found six genetic loci that are associated with recombination phenotypes. Two of these (RNF212 and an inversion on chromosome 17q21.31) were previously reported in the Icelandic population, and this is the first replication in any other population. Of the four newly identified loci (KIAA1462, PDZK1, UGCG, NUB1), results from expression studies provide support for their roles in meiosis. Each of the variants that we identified explains only a small fraction of the individual variation in recombination. Notably, we found different sequence variants associated with female and male recombination phenotypes, suggesting that they are regulated by different genes. Characterization of genetic variants that influence natural variation in meiotic recombination will lead to a better understanding of normal meiotic events as well as of non-disjunction, the primary cause of pregnancy loss.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
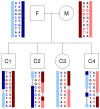
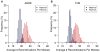
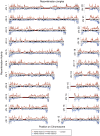
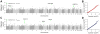

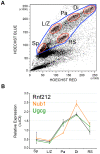
References
-
- Coop G, Wen X, Ober C, Pritchard JK, Przeworski M. High-resolution mapping of crossovers reveals extensive variation in fine-scale recombination patterns among humans. Science. 2008;319:1395–1398. - PubMed
-
- Shifman S, Bell JT, Copley RR, Taylor MS, Williams RW, et al. A high-resolution single nucleotide polymorphism genetic map of the mouse genome. PLoS Biol. 2006;4:e395. doi: 10.1371/journal.pbio.0040395. - DOI - PMC - PubMed
-
- Petkov PM, Broman KW, Szatkiewicz JP, Paigen K. Crossover interference underlies sex differences in recombination rates. Trends Genet. 2007;23:539–542. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

