The tissue-specific expression of TRPML2 (MCOLN-2) gene is influenced by the presence of TRPML1
- PMID: 19763610
- PMCID: PMC2913554
- DOI: 10.1007/s00424-009-0716-5
The tissue-specific expression of TRPML2 (MCOLN-2) gene is influenced by the presence of TRPML1
Abstract
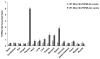
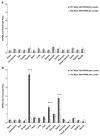
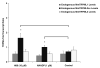
Mucolipidosis type IV is a lysosomal storage disorder caused by the loss or dysfunction of the mucolipin-1 (TRPML1) protein. It has been suggested that TRPML2 could genetically compensate (i.e., become upregulated) for the loss of TRPML1. We thus investigated this possibility by first studying the expression pattern of mouse TRPML2 and its basic channel properties using the varitint-waddler (Va) model. Here, we confirmed the presence of long variant TRPML2 (TRPML2lv) and short variant (TRPML2sv) isoforms. We showed for the first time that, heterologously expressed, TRPML2lv-Va is an active, inwardly rectifying channel. Secondly, we quantitatively measured TRPML2 and TRPML3 mRNA expressions in TRPML1-/- null and wild-type (Wt) mice. In wild-type mice, the TRPML2lv transcripts were very low while TRPML2sv and TRPML3 transcripts have predominant expressions in lymphoid and kidney organs. Significant reductions of TRPML2sv, but not TRPML2lv or TRPML3 transcripts, were observed in lymphoid and kidney organs of TRPML1-/- mice. RNA interference of endogenous human TRPML1 in HEK-293 cells produced a comparable decrease of human TRPML2 transcript levels that can be restored by overexpression of human TRPML1. Conversely, significant upregulation of TRPML2sv transcripts was observed when primary mouse lymphoid cells were treated with nicotinic acid adenine dinucleotide phosphate, or N-(2-[p-bromocinnamylamino]ethyl)-5-isoquinoline sulfonamide, both known activators of TRPML1. In conclusion, our results indicate that TRPML2 is unlikely to compensate for the loss of TRPML1 in lymphoid or kidney organs and that TRPML1 appears to play a novel role in the tissue-specific transcriptional regulation of TRPML2.
Figures



Similar articles
-
Constitutive activity of the human TRPML2 channel induces cell degeneration.J Biol Chem. 2010 Jan 22;285(4):2771-82. doi: 10.1074/jbc.M109.046508. Epub 2009 Nov 23. J Biol Chem. 2010. PMID: 19940139 Free PMC article.
-
Lysosomal localization of TRPML3 depends on TRPML2 and the mucolipidosis-associated protein TRPML1.J Biol Chem. 2006 Jun 23;281(25):17517-17527. doi: 10.1074/jbc.M600807200. Epub 2006 Apr 10. J Biol Chem. 2006. PMID: 16606612 Free PMC article.
-
Mucolipidosis type IV and the mucolipins.Biochem Soc Trans. 2010 Dec;38(6):1432-5. doi: 10.1042/BST0381432. Biochem Soc Trans. 2010. PMID: 21118102
-
The mucolipin-2 (TRPML2) ion channel: a tissue-specific protein crucial to normal cell function.Pflugers Arch. 2016 Feb;468(2):177-92. doi: 10.1007/s00424-015-1732-2. Epub 2015 Sep 4. Pflugers Arch. 2016. PMID: 26336837 Free PMC article. Review.
-
TRPML2 and mucolipin evolution.Handb Exp Pharmacol. 2014;222:647-58. doi: 10.1007/978-3-642-54215-2_25. Handb Exp Pharmacol. 2014. PMID: 24756724 Review.
Cited by
-
Emerging Role of Mucolipins TRPML Channels in Cancer.Front Oncol. 2020 Apr 28;10:659. doi: 10.3389/fonc.2020.00659. eCollection 2020. Front Oncol. 2020. PMID: 32411610 Free PMC article. No abstract available.
-
Mucolipin co-deficiency causes accelerated endolysosomal vacuolation of enterocytes and failure-to-thrive from birth to weaning.PLoS Genet. 2014 Dec 18;10(12):e1004833. doi: 10.1371/journal.pgen.1004833. eCollection 2014 Dec. PLoS Genet. 2014. PMID: 25521295 Free PMC article.
-
Constitutive activity of the human TRPML2 channel induces cell degeneration.J Biol Chem. 2010 Jan 22;285(4):2771-82. doi: 10.1074/jbc.M109.046508. Epub 2009 Nov 23. J Biol Chem. 2010. PMID: 19940139 Free PMC article.
-
Transient receptor potential channels: current perspectives on evolution, structure, function and nomenclature.Proc Biol Sci. 2020 Aug 26;287(1933):20201309. doi: 10.1098/rspb.2020.1309. Epub 2020 Aug 26. Proc Biol Sci. 2020. PMID: 32842926 Free PMC article.
-
NAADP receptors.Cold Spring Harb Perspect Biol. 2011 Jan 1;3(1):a004036. doi: 10.1101/cshperspect.a004036. Cold Spring Harb Perspect Biol. 2011. PMID: 21047915 Free PMC article. Review.
References
-
- Bargal R, Goebel HH, Latta E, Bach G. Mucolipidosis IV: novel mutation and diverse ultrastructural spectrum in the skin. Neuropediatrics. 2002;33:199–202. - PubMed
-
- Sun M, Goldin E, Stahl S, Falardeau JL, Kennedy JC, Acierno JS, Jr, Bove C, Kaneski CR, Nagle J, Bromley MC, Colman M, Schiffmann R, Slaugenhaupt SA. Mucolipidosis type IV is caused by mutations in a gene encoding a novel transient receptor potential channel. Hum Mol Genet. 2000;9:2471–2478. - PubMed
-
- Amir N, Zlotogora J, Bach G. Mucolipidosis type IV: clinical spectrum and natural history. Pediatrics. 1987;79:953–959. - PubMed
-
- Bach G, Cohen MM, Kohn G. Abnormal ganglioside accumulation in cultured fibroblasts from patients with mucolipidosis IV. Biochem Biophys Res Commun. 1975;66:1483–1490. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

