Heterozygous NTF4 mutations impairing neurotrophin-4 signaling in patients with primary open-angle glaucoma
- PMID: 19765683
- PMCID: PMC2756554
- DOI: 10.1016/j.ajhg.2009.08.016
Heterozygous NTF4 mutations impairing neurotrophin-4 signaling in patients with primary open-angle glaucoma
Abstract
Glaucoma, a main cause of blindness in the developed world, is characterized by progressive degeneration of retinal ganglion cells (RGCs), resulting in irreversible loss of vision. Although members of the neurotrophin gene family in various species are known to support the survival of numerous neuronal populations, including RGCs, it is less clear whether they are also required for survival and maintenance of adult neurons in humans. Here, we report seven different heterozygous mutations in the Neurotrophin-4 (NTF4) gene accounting for about 1.7% of primary open-angle glaucoma patients of European origin. Molecular modeling predicted a decreased affinity of neurotrophin 4 protein (NT-4) mutants with its specific tyrosine kinase receptor B (TrkB). Expression of recombinant NT-4 carrying the most frequent mutation was demonstrated to lead to decreased activation of TrkB. These findings suggest a pathway in the pathophysiology of glaucoma through loss of neurotrophic function and may eventually open the possibility of using ligands activating TrkB to prevent the progression of the disease.
Figures
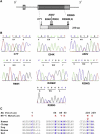




Comment in
-
No evidence of association of heterozygous NTF4 mutations in patients with primary open-angle glaucoma.Am J Hum Genet. 2010 Mar 12;86(3):498-9; author reply 500. doi: 10.1016/j.ajhg.2009.11.018. Am J Hum Genet. 2010. PMID: 20215012 Free PMC article. No abstract available.
References
-
- Tuck M.W., Crick R.P. The projected increase in glaucoma due to an ageing population. Ophthalmic Physiol. Opt. 2003;23:175–179. - PubMed
-
- Nickells R.W. Retinal ganglion cell death in glaucoma: the how, the why, and the maybe. J. Glaucoma. 1996;5:345–356. - PubMed
-
- Quigley H.A. Neuronal death in glaucoma. Prog. Retin. Eye Res. 1999;18:39–57. - PubMed
-
- Soto I., Oglesby E., Buckingham B.P., Son J.L., Roberson E.D., Steele M.R., Inman D.M., Vetter M.L., Horner P.J., Marsh-Armstrong N. Retinal ganglion cells downregulate gene expression and lose their axons within the optic nerve head in a mouse glaucoma model. J. Neurosci. 2008;28:548–561. - PMC - PubMed
-
- Lutjen-Drecoll E., Kruse F.E. Ophthalmologe. 2007;104:167–178. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

