Small effect of fragmentation on the genetic diversity of Dalbergia monticola, an endangered tree species of the eastern forest of Madagascar, detected by chloroplast and nuclear microsatellites
- PMID: 19773273
- PMCID: PMC2766213
- DOI: 10.1093/aob/mcp231
Small effect of fragmentation on the genetic diversity of Dalbergia monticola, an endangered tree species of the eastern forest of Madagascar, detected by chloroplast and nuclear microsatellites
Abstract
Background and aims: The oriental forest ecosystem in Madagascar has been seriously impacted by fragmentation. The pattern of genetic diversity was analysed on a tree species, Dalbergia monticola, which plays an important economic role in Madagascar and is one of the many endangered tree species in the eastern forest.
Methods: Leaves from 546 individuals belonging to 18 small populations affected by different levels of fragmentation were genotyped using eight nuclear (nuc) and three chloroplast (cp) microsatellite markers.
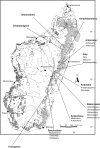
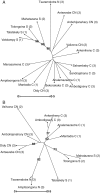
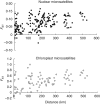
Key results: For nuclear microsatellites, allelic richness (R) and heterozygosity (H(e,nuc)) differed between types of forest: R = 7.36 and R = 9.55, H(e,nuc) = 0.64 and H(e,nuc) = 0.80 in fragmented and non-fragmented forest, respectively, but the differences were not significant. Only the mean number of alleles (N(a,nuc)) and the fixation index F(IS) differed significantly: N(a,nuc) = 9.41 and N(a,nuc) = 13.18, F(IS) = 0.06 and F(IS) = 0.15 in fragmented and non-fragmented forests, respectively. For chloroplast microsatellites, estimated genetic diversity was higher in non-fragmented forest, but the difference was not significant. No recent bottleneck effect was detected for either population. Overall differentiation was low for nuclear microsatellites (F(ST,nuc) = 0.08) and moderate for chloroplast microsatellites (F(ST,cp) = 0.49). A clear relationship was observed between genetic and geographic distance (r = 0.42 P < 0.01 and r = 0.42 P = 0.03 for nuclear and chloroplast microsatellites, respectively), suggesting a pattern of isolation by distance. Analysis of population structure using the neighbor-joining method or Bayesian models separated southern populations from central and northern populations with nuclear microsatellites, and grouped the population according to regions with chloroplast microsatellites, but did not separate the fragmented populations.
Conclusions: Residual diversity and genetic structure of populations of D. monticola in Madagascar suggest a limited impact of fragmentation on molecular genetic parameters.
Figures
Similar articles
-
Genetic diversity and gene flow in a Caribbean tree Pterocarpus officinalis Jacq.: a study based on chloroplast and nuclear microsatellites.Genetica. 2009 Mar;135(2):185-98. doi: 10.1007/s10709-008-9268-4. Epub 2008 Apr 23. Genetica. 2009. PMID: 18431679
-
Diversity and genetic connectivity among populations of a threatened tree (Dalbergia nigra) in a recently fragmented landscape of the Brazilian Atlantic Forest.Genetica. 2011 Sep;139(9):1159-68. doi: 10.1007/s10709-011-9618-5. Epub 2011 Nov 30. Genetica. 2011. PMID: 22127549
-
Habitat loss other than fragmentation per se decreased nuclear and chloroplast genetic diversity in a monoecious tree.PLoS One. 2012;7(6):e39146. doi: 10.1371/journal.pone.0039146. Epub 2012 Jun 18. PLoS One. 2012. PMID: 22723951 Free PMC article.
-
Long-distance seed and pollen dispersal inferred from spatial genetic structure in the very low-density rainforest tree, Baillonella toxisperma Pierre, in Central Africa.Mol Ecol. 2010 Nov;19(22):4949-62. doi: 10.1111/j.1365-294X.2010.04864.x. Epub 2010 Oct 21. Mol Ecol. 2010. PMID: 20964756
-
A novel synthesis of two decades of microsatellite studies on European beech reveals decreasing genetic diversity from glacial refugia.Tree Genet Genomes. 2023;19(1):3. doi: 10.1007/s11295-022-01577-4. Epub 2022 Dec 12. Tree Genet Genomes. 2023. PMID: 36532711 Free PMC article. Review.
Cited by
-
Molecular markers from the chloroplast genome of rose provide a complementary tool for variety discrimination and profiling.Sci Rep. 2020 Jul 22;10(1):12188. doi: 10.1038/s41598-020-68092-1. Sci Rep. 2020. PMID: 32699274 Free PMC article.
-
Spatial patterns of AFLP diversity in Bulbophyllum occultum (Orchidaceae) indicate long-term refugial isolation in Madagascar and long-distance colonization effects in La Réunion.Heredity (Edinb). 2016 May;116(5):434-46. doi: 10.1038/hdy.2016.1. Epub 2016 Feb 17. Heredity (Edinb). 2016. PMID: 26883184 Free PMC article.
-
Genetic structure and diversity of natural and domesticated populations of Citrus medica L. in the Eastern Himalayan region of Northeast India.Ecol Evol. 2016 May 10;6(12):3898-911. doi: 10.1002/ece3.2174. eCollection 2016 Jun. Ecol Evol. 2016. PMID: 27516853 Free PMC article.
-
Population genetic structure of the endemic rosewoods Dalbergia cochinchinensis and D. oliveri at a regional scale reflects the Indochinese landscape and life-history traits.Ecol Evol. 2017 Dec 1;8(1):530-545. doi: 10.1002/ece3.3626. eCollection 2018 Jan. Ecol Evol. 2017. PMID: 29321891 Free PMC article.
-
Drivers of population divergence and species differentiation in a recent group of indigenous orchids (Vanilla spp.) in Madagascar.Ecol Evol. 2021 Feb 24;11(6):2681-2700. doi: 10.1002/ece3.7224. eCollection 2021 Mar. Ecol Evol. 2021. PMID: 33767829 Free PMC article.
References
-
- Addinsoft. XLSTAT software version 7.5.2. 2008. http://www.xlstat.com. (accessed 13 February 2009)
-
- Aguilar R, Quesada M, Ashworth L, Herrerias-Diego Y, Lobo J. Genetic consequences of habitat fragmentation in plant populations: susceptible signals in plant traits and methodological approaches. Molecular Ecology. 2008;17:5177–5188. - PubMed
-
- Aldrich PR, Hamrick JL, Chavarriaga P, Kochert G. Microsatellite analysis of demographic genetic structure in fragmented populations of the tropical tree Symphonia globulifera. Molecular Ecology. 1998;7:933–944. - PubMed
-
- Andre T, Lemes MR, Grogan J, Gribel R. Post-logging loss of genetic diversity in a mahogany (Swietenia macrophylla King, Meliaceae) population in Brazilian Amazonia. Forest Ecology and Management. 2008;255:340–345.
-
- Andrianoelina O, Rakotondraoelina H, Ramamonjisoa L, Maley J, Danthu P, Bouvet J-M. Genetic diversity of Dalbergia monticola (Fabaceae) an endangered species in the fragmented oriental forest of Madagascar. Biodiversity Conservation. 2006;15:1109–1128.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

