Imprinting of the polycomb group gene MEDEA serves as a ploidy sensor in Arabidopsis
- PMID: 19779546
- PMCID: PMC2738949
- DOI: 10.1371/journal.pgen.1000663
Imprinting of the polycomb group gene MEDEA serves as a ploidy sensor in Arabidopsis
Abstract
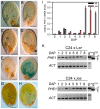
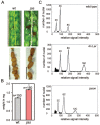
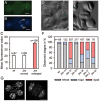
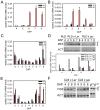
Balanced maternal and paternal genome contributions are a requirement for successful seed development. Unbalanced contributions often cause seed abortion, a phenomenon that has been termed "triploid block." Misregulation of imprinted regulatory genes has been proposed to be the underlying cause for abnormalities in growth and structure of the endosperm in seeds with deviating parental contributions. We identified a mutant forming unreduced pollen that enabled us to investigate direct effects of unbalanced parental genome contributions on seed development and to reveal the underlying molecular mechanism of dosage sensitivity. We provide evidence that parent-of-origin-specific expression of the Polycomb group (PcG) gene MEDEA is causally responsible for seed developmental aberrations in Arabidopsis seeds with increased paternal genome contributions. We propose that imprinted expression of PcG genes is an evolutionary conserved mechanism to balance parental genome contributions in embryo nourishing tissues.
Conflict of interest statement
European patent EP09008196 “Polyploid plants” was deposited by ETH on June 23rd, 2009.
Figures



Similar articles
-
Parental genome dosage imbalance deregulates imprinting in Arabidopsis.PLoS Genet. 2010 Mar 19;6(3):e1000885. doi: 10.1371/journal.pgen.1000885. PLoS Genet. 2010. PMID: 20333248 Free PMC article.
-
The Polycomb-group protein MEDEA regulates seed development by controlling expression of the MADS-box gene PHERES1.Genes Dev. 2003 Jun 15;17(12):1540-53. doi: 10.1101/gad.257403. Genes Dev. 2003. PMID: 12815071 Free PMC article.
-
Imprinting of the MEDEA polycomb gene in the Arabidopsis endosperm.Plant Cell. 1999 Oct;11(10):1945-52. doi: 10.1105/tpc.11.10.1945. Plant Cell. 1999. PMID: 10521524 Free PMC article.
-
[Genomic imprinting and seed development].Yi Chuan. 2005 Jul;27(4):665-70. Yi Chuan. 2005. PMID: 16120596 Review. Chinese.
-
Epigenetic mechanisms governing seed development in plants.EMBO Rep. 2006 Dec;7(12):1223-7. doi: 10.1038/sj.embor.7400854. EMBO Rep. 2006. PMID: 17139298 Free PMC article. Review.
Cited by
-
Parental genome dosage imbalance deregulates imprinting in Arabidopsis.PLoS Genet. 2010 Mar 19;6(3):e1000885. doi: 10.1371/journal.pgen.1000885. PLoS Genet. 2010. PMID: 20333248 Free PMC article.
-
Evolutionary studies of the bHLH transcription factors belonging to MBW complex: their role in seed development.Ann Bot. 2023 Nov 23;132(3):383-400. doi: 10.1093/aob/mcad097. Ann Bot. 2023. PMID: 37467144 Free PMC article. Review.
-
The emerging importance of type I MADS box transcription factors for plant reproduction.Plant Cell. 2011 Mar;23(3):865-72. doi: 10.1105/tpc.110.081737. Epub 2011 Mar 4. Plant Cell. 2011. PMID: 21378131 Free PMC article. Review.
-
Unveiling the imprinted dance: how parental genomes orchestrate seed development and hybrid success.Front Plant Sci. 2024 Sep 27;15:1455685. doi: 10.3389/fpls.2024.1455685. eCollection 2024. Front Plant Sci. 2024. PMID: 39399543 Free PMC article. Review.
-
Molecular tools for exploring polyploid genomes in plants.Int J Mol Sci. 2012;13(8):10316-10335. doi: 10.3390/ijms130810316. Epub 2012 Aug 17. Int J Mol Sci. 2012. PMID: 22949863 Free PMC article. Review.
References
-
- Otto SP, Whitton J. Polyploid incidence and evolution. Annu Rev Genet. 2000;34:401–437. - PubMed
-
- Comai L. The advantages and disadvantages of being polyploid. Nat Rev Genet. 2005;6:836–846. - PubMed
-
- Hegarty MJ, Hiscock SJ. Genomic clues to the evolutionary success of polyploid plants. Curr Biol. 2008;18:R435–44. - PubMed
-
- Ramsey S, Schemske D. Pathways, Mechanisms, and rates of polyploid formation in flowering plants. Auual Reviews Ecology and Systematics. 1998;29:467–501.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

