Novel approaches to inhibiting HIV-1 replication
- PMID: 19782103
- PMCID: PMC2815006
- DOI: 10.1016/j.antiviral.2009.09.009
Novel approaches to inhibiting HIV-1 replication
Abstract
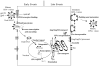
Considerable success has been achieved in the treatment of HIV-1 infection, and more than two-dozen antiretroviral drugs are available targeting several distinct steps in the viral replication cycle. However, resistance to these compounds emerges readily, even in the context of combination therapy. Drug toxicity, adverse drug-drug interactions, and accompanying poor patient adherence can also lead to treatment failure. These considerations make continued development of novel antiretroviral therapeutics necessary. In this article, we highlight a number of steps in the HIV-1 replication cycle that represent promising targets for drug discovery. These include lipid raft microdomains, the RNase H activity of the viral enzyme reverse transcriptase, uncoating of the viral core, host cell machinery involved in the integration of the viral DNA into host cell chromatin, virus assembly, maturation, and budding, and the functions of several viral accessory proteins. We discuss the relevant molecular and cell biology, and describe progress to date in developing inhibitors against these novel targets. This article forms part of a special issue of Antiviral Research marking the 25th anniversary of antiretroviral drug discovery and development, Vol 85, issue 1, 2010.
Published by Elsevier B.V.
Figures


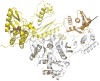









References
-
- Abdurahman S, Hoglund S, Goobar-Larsson L, Vahlne A. Selected amino acid substitutions in the C-terminal region of human immunodeficiency virus type 1 capsid protein affect virus assembly and release. J Gen Virol. 2004;85(Pt 10):2903–13. - PubMed
-
- Adamson CS, Freed EO. HIV-1 Assembly, Release and Maturation. In: Jeang KT, editor. Advances in Pharmacolgy, HIV-1: Molecular Biology and Pathogenesis: Viral Mechansims. Vol. 55. Elsevier; 2007.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

