Involvement of the beta clamp in methyl-directed mismatch repair in vitro
- PMID: 19783657
- PMCID: PMC2781695
- DOI: 10.1074/jbc.M109.054528
Involvement of the beta clamp in methyl-directed mismatch repair in vitro
Abstract
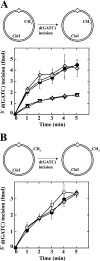
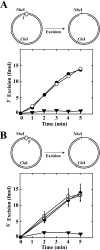
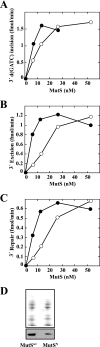
We have examined function of the bacterial beta replication clamp in the different steps of methyl-directed DNA mismatch repair. The mismatch-, MutS-, and MutL-dependent activation of MutH is unaffected by the presence or orientation of loaded beta clamp on either 3' or 5' heteroduplexes. Similarly, beta is not required for 3' or 5' mismatch-provoked excision when scored in the presence of gamma complex or in the presence of gamma complex and DNA polymerase III core components. However, mismatch repair does not occur in the absence of beta, an effect we attribute to a requirement for the clamp in the repair DNA synthesis step of the reaction. We have confirmed previous findings that beta clamp interacts specifically with MutS and MutL (López de Saro, F. J., Marinus, M. G., Modrich, P., and O'Donnell, M. (2006) J. Biol. Chem. 281, 14340-14349) and show that the mutator phenotype conferred by amino acid substitution within the MutS N-terminal beta-interaction motif is the probable result of instability coupled with reduced activity in multiple steps of the repair reaction. In addition, we have found that the DNA polymerase III alpha catalytic subunit interacts strongly and specifically with both MutS and MutL. Because interactions of polymerase III holoenzyme components with MutS and MutL appear to be of limited import during the initiation and excision steps of mismatch correction, we suggest that their significance might lie in the control of replication fork events in response to the sensing of DNA lesions by the repair system.
Figures





References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

