Nucleotide excision repair-induced H2A ubiquitination is dependent on MDC1 and RNF8 and reveals a universal DNA damage response
- PMID: 19797077
- PMCID: PMC2753161
- DOI: 10.1083/jcb.200902150
Nucleotide excision repair-induced H2A ubiquitination is dependent on MDC1 and RNF8 and reveals a universal DNA damage response
Abstract
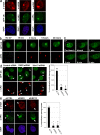
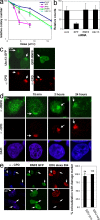
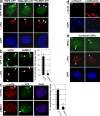
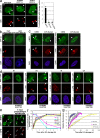
Chromatin modifications are an important component of the of DNA damage response (DDR) network that safeguard genomic integrity. Recently, we demonstrated nucleotide excision repair (NER)-dependent histone H2A ubiquitination at sites of ultraviolet (UV)-induced DNA damage. In this study, we show a sustained H2A ubiquitination at damaged DNA, which requires dynamic ubiquitination by Ubc13 and RNF8. Depletion of these enzymes causes UV hypersensitivity without affecting NER, which is indicative of a function for Ubc13 and RNF8 in the downstream UV-DDR. RNF8 is targeted to damaged DNA through an interaction with the double-strand break (DSB)-DDR scaffold protein MDC1, establishing a novel function for MDC1. RNF8 is recruited to sites of UV damage in a cell cycle-independent fashion that requires NER-generated, single-stranded repair intermediates and ataxia telangiectasia-mutated and Rad3-related protein. Our results reveal a conserved pathway of DNA damage-induced H2A ubiquitination for both DSBs and UV lesions, including the recruitment of 53BP1 and Brca1. Although both lesions are processed by independent repair pathways and trigger signaling responses by distinct kinases, they eventually generate the same epigenetic mark, possibly functioning in DNA damage signal amplification.
Figures

References
-
- Abraham R.T. 2004. PI 3-kinase related kinases: ‘big’ players in stress-induced signaling pathways.DNA Repair (Amst.). 3:883–887 doi:10.1016/j.dnarep.2004.04.002 - DOI - PubMed
-
- Baarends W.M., Wassenaar E., van der Laan R., Hoogerbrugge J., Sleddens-Linkels E., Hoeijmakers J.H., de Boer P., Grootegoed J.A. 2005. Silencing of unpaired chromatin and histone H2A ubiquitination in mammalian meiosis.Mol. Cell. Biol. 25:1041–1053 doi:10.1128/MCB.25.3.1041-1053.2005 - DOI - PMC - PubMed
-
- Bekker-Jensen S., Lukas C., Kitagawa R., Melander F., Kastan M.B., Bartek J., Lukas J. 2006. Spatial organization of the mammalian genome surveillance machinery in response to DNA strand breaks.J. Cell Biol. 173:195–206 doi:10.1083/jcb.200510130 - DOI - PMC - PubMed
-
- Bennett E.J., Harper J.W. 2008. DNA damage: ubiquitin marks the spot.Nat. Struct. Mol. Biol. 15:20–22 doi:10.1038/nsmb0108-20 - DOI - PubMed
-
- Bergink S., Salomons F.A., Hoogstraten D., Groothuis T.A., de Waard H., Wu J., Yuan L., Citterio E., Houtsmuller A.B., Neefjes J., et al. 2006. DNA damage triggers nucleotide excision repair-dependent monoubiquitylation of histone H2A.Genes Dev. 20:1343–1352 doi:10.1101/gad.373706 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

