Genome-wide comparative analysis of the Brassica rapa gene space reveals genome shrinkage and differential loss of duplicated genes after whole genome triplication
- PMID: 19821981
- PMCID: PMC2784326
- DOI: 10.1186/gb-2009-10-10-r111
Genome-wide comparative analysis of the Brassica rapa gene space reveals genome shrinkage and differential loss of duplicated genes after whole genome triplication
Abstract
Background: Brassica rapa is one of the most economically important vegetable crops worldwide. Owing to its agronomic importance and phylogenetic position, B. rapa provides a crucial reference to understand polyploidy-related crop genome evolution. The high degree of sequence identity and remarkably conserved genome structure between Arabidopsis and Brassica genomes enables comparative tiling sequencing using Arabidopsis sequences as references to select the counterpart regions in B. rapa, which is a strong challenge of structural and comparative crop genomics.
Results: We assembled 65.8 megabase-pairs of non-redundant euchromatic sequence of B. rapa and compared this sequence to the Arabidopsis genome to investigate chromosomal relationships, macrosynteny blocks, and microsynteny within blocks. The triplicated B. rapa genome contains only approximately twice the number of genes as in Arabidopsis because of genome shrinkage. Genome comparisons suggest that B. rapa has a distinct organization of ancestral genome blocks as a result of recent whole genome triplication followed by a unique diploidization process. A lack of the most recent whole genome duplication (3R) event in the B. rapa genome, atypical of other Brassica genomes, may account for the emergence of B. rapa from the Brassica progenitor around 8 million years ago.
Conclusions: This work demonstrates the potential of using comparative tiling sequencing for genome analysis of crop species. Based on a comparative analysis of the B. rapa sequences and the Arabidopsis genome, it appears that polyploidy and chromosomal diploidization are ongoing processes that collectively stabilize the B. rapa genome and facilitate its evolution.
Figures



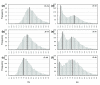


References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

