Diversity of the Arabidopsis mitochondrial genome occurs via nuclear-controlled recombination activity
- PMID: 19822729
- PMCID: PMC2787419
- DOI: 10.1534/genetics.109.108514
Diversity of the Arabidopsis mitochondrial genome occurs via nuclear-controlled recombination activity
Abstract
The plant mitochondrial genome is recombinogenic, with DNA exchange activity controlled to a large extent by nuclear gene products. One nuclear gene, MSH1, appears to participate in suppressing recombination in Arabidopsis at every repeated sequence ranging in size from 108 to 556 bp. Present in a wide range of plant species, these mitochondrial repeats display evidence of successful asymmetric DNA exchange in Arabidopsis when MSH1 is disrupted. Recombination frequency appears to be influenced by repeat sequence homology and size, with larger size repeats corresponding to increased DNA exchange activity. The extensive mitochondrial genomic reorganization of the msh1 mutant produced altered mitochondrial transcription patterns. Comparison of mitochondrial genomes from the Arabidopsis ecotypes C24, Col-0, and Ler suggests that MSH1 activity accounts for most or all of the polymorphisms distinguishing these genomes, producing ecotype-specific stoichiometric changes in each line. Our observations suggest that MSH1 participates in mitochondrial genome evolution by influencing the lineage-specific pattern of mitochondrial genetic variation in higher plants.
Figures



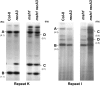

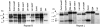


References
-
- Belliard, G., F. Vedel and G. Pelletier, 1979. Mitochondrial recombination in cytoplasmic hybrids of Nicotiana tabacum by protoplast fusion. Nature 281 401–403.
-
- Feng, X., A. P. Kaur, S. A. Mackenzie and I. M. Dweikat, 2009. Substoichiometric shifting in the fertility reversion of cytoplasmic male sterile pearl millet. Theor. Appl. Genet. 118 1361–1370. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

