Structural biology of copper trafficking
- PMID: 19824702
- PMCID: PMC2768115
- DOI: 10.1021/cr900104z
Structural biology of copper trafficking
Figures



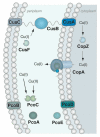
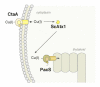


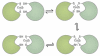

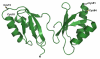





References
-
- Crichton RR, Pierre J-L. Biometals. 2001;14:99. - PubMed
-
- da Silva J. J. R. Fraústo, Williams RJP. The biological chemistry of the elements. Second ed Oxford University Press; Oxford: 2001.
-
- Kaim W, Rall J. Angew. Chem. Int. Ed. Engl. 1996;35:43.
-
- Ochiai EI. J. Chem. Ed. 1986;63:942.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

