Minimal membrane docking requirements revealed by reconstitution of Rab GTPase-dependent membrane fusion from purified components
- PMID: 19826089
- PMCID: PMC2764952
- DOI: 10.1073/pnas.0903801106
Minimal membrane docking requirements revealed by reconstitution of Rab GTPase-dependent membrane fusion from purified components
Abstract
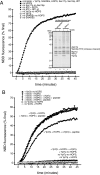
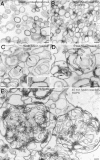
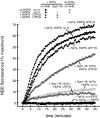
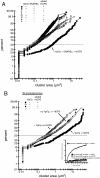
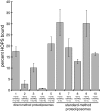
Rab GTPases and their effectors mediate docking, the initial contact of intracellular membranes preceding bilayer fusion. However, it has been unclear whether Rab proteins and effectors are sufficient for intermembrane interactions. We have recently reported reconstituted membrane fusion that requires yeast vacuolar SNAREs, lipids, and the homotypic fusion and vacuole protein sorting (HOPS)/class C Vps complex, an effector and guanine nucleotide exchange factor for the yeast vacuolar Rab GTPase Ypt7p. We now report reconstitution of lysis-free membrane fusion that requires purified GTP-bound Ypt7p, HOPS complex, vacuolar SNAREs, ATP hydrolysis, and the SNARE disassembly catalysts Sec17p and Sec18p. We use this reconstituted system to show that SNAREs and Sec17p/Sec18p, and Ypt7p and the HOPS complex, are required for stable intermembrane interactions and that the three vacuolar Q-SNAREs are sufficient for these interactions.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Comment in
-
Organelle docking: R-SNAREs are late.Proc Natl Acad Sci U S A. 2009 Nov 24;106(47):19745-6. doi: 10.1073/pnas.0911425106. Epub 2009 Nov 18. Proc Natl Acad Sci U S A. 2009. PMID: 19923425 Free PMC article. No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

