Structural perturbations induced by the alpha-anomer of the aflatoxin B(1) formamidopyrimidine adduct in duplex and single-strand DNA
- PMID: 19831353
- PMCID: PMC2773149
- DOI: 10.1021/ja902052v
Structural perturbations induced by the alpha-anomer of the aflatoxin B(1) formamidopyrimidine adduct in duplex and single-strand DNA
Abstract
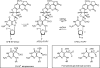
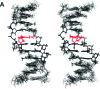
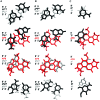
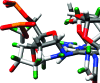
The guanine N7 adduct of aflatoxin B(1) exo-8,9-epoxide hydrolyzes to form the formamidopyrimidine (AFB-FAPY) adduct, which interconverts between alpha and beta anomers. The beta anomer is highly mutagenic in Escherichia coli, producing G --> T transversions; it thermally stabilizes the DNA duplex. The AFB-alpha-FAPY adduct blocks replication; it destabilizes the DNA duplex. Herein, the structure of the AFB-alpha-FAPY adduct has been elucidated in 5'-d(C(1)T(2)A(3)T(4)X(5)A(6)T(7)T(8)C(9)A(10))-3'.5'-d(T(11)G(12)A(13)A(14)T(15)C(16)A(17)T(18)A(19)G(20))-3' (X = AFB-alpha-FAPY) using molecular dynamics calculations restrained by NMR-derived distances and torsion angles. The AFB moiety intercalates on the 5' face of the pyrimidine moiety at the damaged nucleotide between base pairs T(4).A(17) and X(5).C(16), placing the FAPY C5-N(5) bond in the R(a) axial conformation. Large perturbations of the epsilon and zeta backbone torsion angles are observed, and the base stacking register of the duplex is perturbed. The deoxyribose orientation shifts to become parallel to the FAPY base and displaced toward the minor groove. Intrastrand stacking between the AFB moiety and the 5' neighbor thymine remains, but strong interstrand stacking is not observed. A hydrogen bond between the formyl group and the exocyclic amine of the 3'-neighbor adenine stabilizes the E conformation of the formamide moiety. NMR studies reveal a similar 5'-intercalation of the AFB moiety for the AFB-alpha-FAPY adduct in the tetramer 5'-d(C(1)T(2)X(3)A(4))-3', involving the R(a) axial conformation of the FAPY C5-N(5) bond and the E conformation of the formamide moiety. Since in duplex DNA the AFB moiety of the AFB-beta-FAPY adduct also intercalates on the 5' side of the pyrimidine moiety at the damaged nucleotide, we conclude that favorable 5'-stacking leads to the R(a) conformational preference about the C5-N(5) bond; the same conformational preference about this bond is also observed at the nucleoside and base levels. The structural distortions and the less favorable stacking interactions induced by the AFB-alpha-FAPY adduct explain its lower stability as compared to the AFB-beta-FAPY adduct in duplex DNA. In this DNA sequence, hydrogen bonding between the formyl oxygen and the exocyclic amine of the 3'-neighboring adenine stabilizing the E configuration of the formamide moiety is also observed for the AFB-beta-FAPY adduct, and suggests that the identity of the 3'-neighbor nucleotide modulates the stability and biological processing of AFB adducts.
Figures




References
-
- Busby W. F. Jr.; Wogan G. N. In. Chemical Carcinogens; 2nd ed.; Searle C. E., Ed.; American Chemical Society: Washington, D.C., 1984; pp 945−1136.
-
- Smela M. E.; Currier S. S.; Bailey E. A.; Essigmann J. M. Carcinogenesis 2001, 22, 535–545. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

