Allelic inactivation of rDNA loci
- PMID: 19833769
- PMCID: PMC2764490
- DOI: 10.1101/gad.544509
Allelic inactivation of rDNA loci
Abstract
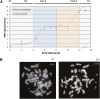
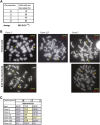
Human cells contain several hundred ribosomal genes (rDNA) that are clustered into nucleolar organizer regions (NORs) on the short arms of five different acrocentric chromosomes. Only approximately 50% of the gene copies are actually expressed in somatic cells. Here, we used a new cytological technique to demonstrate that rDNA is regulated allelically in a regional manner, with one parental copy of each NOR being repressed in any individual cell. This process is similar to that of X-chromosome inactivation in females. Early in development, one copy of each NOR becomes late-replicating, thus probably marking it for inactivation and subsequent targeted de novo methylation at rDNA promoter regions. Once established, this multichromosomal allelic pattern is then maintained clonally in somatic cells. This pathway may serve as an epigenetic mechanism for controlling the number of available rDNA copies during development.
Figures




References
-
- Azuara V, Brown KE, Williams RR, Webb N, Dillon N, Festenstein R, Buckle V, Merkenschlager M, Fisher AG. Heritable gene silencing in lymphocytes delays chromatid resolution without affecting the timing of DNA replication. Nat Cell Biol. 2003;5:668–674. - PubMed
-
- Birger Y, Shemer R, Perk J, Razin A. The imprinting box of the mouse Igf2r gene. Nature. 1999;397:84–88. - PubMed
-
- Boisvert FM, van Koningsbruggen S, Navascues J, Lamond AI. The multifunctional nucleolus. Nat Rev Mol Cell Biol. 2007;8:574–585. - PubMed
-
- Brown DD, Dawid IB. Specific gene amplification in oocytes. Oocyte nuclei contain extrachromosomal replicas of the genes for ribosomal RNA. Science. 1968;160:272–280. - PubMed
-
- Chess A, Simon I, Cedar H, Axel R. Allelic inactivation regulates olfactory receptor gene expression. Cell. 1994;78:823–834. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
