Genetic identification of nascent peptides that induce ribosome stalling
- PMID: 19840930
- PMCID: PMC2787343
- DOI: 10.1074/jbc.M109.039040
Genetic identification of nascent peptides that induce ribosome stalling
Abstract
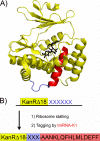
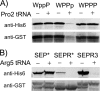
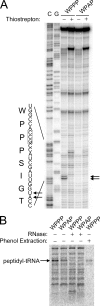
Several nascent peptides stall ribosomes during their own translation in both prokaryotes and eukaryotes. Leader peptides that induce stalling can regulate downstream gene expression. Interestingly, stalling peptides show little sequence similarity and interact with the ribosome through distinct mechanisms. To explore the scope of regulation by stalling peptides and to better understand the mechanism of stalling, we identified and characterized new examples from random libraries. We created a genetic selection that ties the life of Escherichia coli cells to stalling at a specific site. This selection relies on the natural bacterial system that rescues arrested ribosomes. We altered transfer-messenger RNA, a key component of this rescue system, to direct the completion of a necessary protein if and only if stalling occurs. We identified three classes of stalling peptides: C-terminal Pro residues, SecM-like peptides, and the novel stalling sequence FXXYXIWPP. Like the leader peptides SecM and TnaC, the FXXYXIWPP peptide induces stalling efficiently by inhibiting peptidyl transfer. The nascent peptide exit tunnel and peptidyltransferase center are implicated in this stalling event, although mutations in the ribosome affect stalling on SecM and FXXYXIWPP differently. We conclude that ribosome stalling can be caused by numerous sequences and is more common than previously believed.
Figures
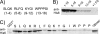

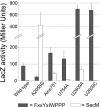
Similar articles
-
Nascent peptides that block protein synthesis in bacteria.Proc Natl Acad Sci U S A. 2013 Mar 5;110(10):E878-87. doi: 10.1073/pnas.1219536110. Epub 2013 Feb 19. Proc Natl Acad Sci U S A. 2013. PMID: 23431150 Free PMC article.
-
Ribosome regulation by the nascent peptide.Microbiol Rev. 1996 Jun;60(2):366-85. doi: 10.1128/mr.60.2.366-385.1996. Microbiol Rev. 1996. PMID: 8801438 Free PMC article. Review.
-
Computational analysis of nascent peptides that induce ribosome stalling and their proteomic distribution in Saccharomyces cerevisiae.RNA. 2017 Jul;23(7):983-994. doi: 10.1261/rna.059188.116. Epub 2017 Mar 31. RNA. 2017. PMID: 28363900 Free PMC article.
-
Mechanisms of ribosome stalling by SecM at multiple elongation steps.Elife. 2015 Dec 14;4:e09684. doi: 10.7554/eLife.09684. Elife. 2015. PMID: 26670735 Free PMC article.
-
Nascent polypeptide sequences that influence ribosome function.Curr Opin Microbiol. 2011 Apr;14(2):160-6. doi: 10.1016/j.mib.2011.01.011. Epub 2011 Feb 20. Curr Opin Microbiol. 2011. PMID: 21342782 Review.
Cited by
-
Amino acid sequence repertoire of the bacterial proteome and the occurrence of untranslatable sequences.Proc Natl Acad Sci U S A. 2016 Jun 28;113(26):7166-70. doi: 10.1073/pnas.1606518113. Epub 2016 Jun 15. Proc Natl Acad Sci U S A. 2016. PMID: 27307442 Free PMC article.
-
RNA-seq reveals multifaceted gene expression response to Fab production in Escherichia coli fed-batch processes with particular focus on ribosome stalling.Microb Cell Fact. 2024 Jan 5;23(1):14. doi: 10.1186/s12934-023-02278-w. Microb Cell Fact. 2024. PMID: 38183013 Free PMC article.
-
Peptidyl transferase center decompaction and structural constraints during early protein elongation on the ribosome.Sci Rep. 2021 Dec 15;11(1):24061. doi: 10.1038/s41598-021-02985-7. Sci Rep. 2021. PMID: 34911999 Free PMC article.
-
Patchy and widespread distribution of bacterial translation arrest peptides associated with the protein localization machinery.Nat Commun. 2024 Apr 2;15(1):2711. doi: 10.1038/s41467-024-46993-3. Nat Commun. 2024. PMID: 38565864 Free PMC article.
-
The ABCF proteins in Escherichia coli individually cope with 'hard-to-translate' nascent peptide sequences.Nucleic Acids Res. 2024 Jun 10;52(10):5825-5840. doi: 10.1093/nar/gkae309. Nucleic Acids Res. 2024. PMID: 38661232 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

