REBASE--a database for DNA restriction and modification: enzymes, genes and genomes
- PMID: 19846593
- PMCID: PMC2808884
- DOI: 10.1093/nar/gkp874
REBASE--a database for DNA restriction and modification: enzymes, genes and genomes
Abstract
REBASE is a comprehensive database of information about restriction enzymes, DNA methyltransferases and related proteins involved in the biological process of restriction-modification (R-M). It contains fully referenced information about recognition and cleavage sites, isoschizomers, neoschizomers, commercial availability, methylation sensitivity, crystal and sequence data. Experimentally characterized homing endonucleases are also included. The fastest growing segment of REBASE contains the putative R-M systems found in the sequence databases. Comprehensive descriptions of the R-M content of all fully sequenced genomes are available including summary schematics. The contents of REBASE may be browsed from the web (http://rebase.neb.com) and selected compilations can be downloaded by ftp (ftp.neb.com). Additionally, monthly updates can be requested via email.
Figures
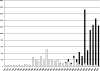
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

