Meta-analysis of genome-wide association studies: no efficiency gain in using individual participant data
- PMID: 19847795
- PMCID: PMC3878085
- DOI: 10.1002/gepi.20435
Meta-analysis of genome-wide association studies: no efficiency gain in using individual participant data
Abstract
To identify genetic variants with modest effects on complex human diseases, a growing number of networks or consortia are created for sharing data from multiple genome-wide association studies on the same disease or related disorders. A central question in this enterprise is whether to obtain summary results or individual participant data from relevant studies. We show theoretically and numerically that meta-analysis of summary results is statistically as efficient as joint analysis of individual participant data (provided that both analyses are performed properly under the same modeling assumptions). We illustrate this equivalence with case-control data from the Finland-United States Investigation of NIDDM Genetics (FUSION) study. Collating only summary results will increase the number and representativeness of available studies, simplify data collection and analysis, reduce resource utilization, and accelerate discovery.
Figures
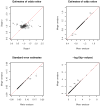
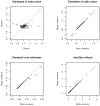
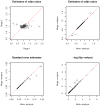
References
-
- Cox DR, Hinkley DV. Theoretical Statistics. Chapman and Hall; 1979.
-
- Kavvoura1 FK, Ioannidis JPA. Methods for meta-analysis in genetic association studies: a review of their potential and pitfalls. Human Genetics. 2008;123:1–14. - PubMed
-
- Olkin I, Sampson A. Comparison of meta-analysis versus analysis of variance of individual patient data. Biometrics. 1998;54:317–22. - PubMed
-
- Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D. Principal components analysis corrects for stratification in genome-wide association studies. Nature Genetics. 2006;38:904–909. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

