Atrogin-1 and MuRF1 regulate cardiac MyBP-C levels via different mechanisms
- PMID: 19850579
- PMCID: PMC4023316
- DOI: 10.1093/cvr/cvp348
Atrogin-1 and MuRF1 regulate cardiac MyBP-C levels via different mechanisms
Abstract
Aims: Familial hypertrophic cardiomyopathy (FHC) is frequently caused by cardiac myosin-binding protein C (cMyBP-C) gene mutations, which should result in C-terminal truncated mutants. However, truncated mutants were not detected in myocardial tissue of FHC patients and were rapidly degraded by the ubiquitin-proteasome system (UPS) after gene transfer in cardiac myocytes. Since the diversity and specificity of UPS regulation lie in E3 ubiquitin ligases, we investigated whether the muscle-specific E3 ligases atrogin-1 or muscle ring finger protein-1 (MuRF1) mediate degradation of truncated cMyBP-C.
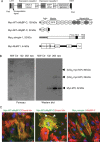
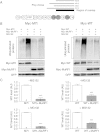
Methods and results: Human wild-type (WT) and truncated (M7t, resulting from a human mutation) cMyBP-C species were co-immunoprecipitated with atrogin-1 after adenoviral overexpression in cardiac myocytes, and WT-cMyBP-C was identified as an interaction partner of MuRF1 by yeast two-hybrid screens. Overexpression of atrogin-1 in cardiac myocytes decreased the protein level of M7t-cMyBP-C by 80% and left WT-cMyBP-C level unaffected. This was rescued by proteasome inhibition. In contrast, overexpression of MuRF1 in cardiac myocytes not only reduced the protein level of WT- and M7t-cMyBP-C by >60%, but also the level of myosin heavy chains (MHCs) by >40%, which were not rescued by proteasome inhibition. Both exogenous cMyBP-C and endogenous MHC mRNA levels were markedly reduced by MuRF1 overexpression. Similar to cardiac myocytes, MuRF1-overexpressing (TG) mice exhibited 40% lower levels of MHC mRNAs and proteins. Protein levels of cMyBP-C were 29% higher in MuRF1 knockout and 34% lower in TG than in WT, without a corresponding change in mRNA levels.
Conclusion: These data suggest that atrogin-1 specifically targets truncated M7t-cMyBP-C, but not WT-cMyBP-C, for proteasomal degradation and that MuRF1 indirectly reduces cMyBP-C levels by regulating the transcription of MHC.
Figures




References
-
- Richardson P, McKenna W, Bristow M, Maish B, Mautner B, O'Connell J, et al. Report of the 1995 World Health Organisation/International Society and Federation of Cardiology task force on the definition and classification of cardiomyopathies. Circulation. 1996;93:841–842. - PubMed
-
- Richard P, Villard E, Charron P, Isnard R. The genetic bases of cardiomyopathies. J Am Coll Cardiol. 2006;48:A79–A89.
-
- Friedrich F, Bausero P, Sun Y, Treszl A, Krämer E, Juhr D, et al. A new polymorphism in human calmodulin III promoter is a potential modifier gene for familial hypertrophic cardiomyopathy. Eur Heart J. 2009;30:1648–1655. - PubMed
-
- Richard P, Charron P, Carrier L, Ledeuil C, Cheav T, Pichereau C, et al. Hypertrophic cardiomyopathy: distribution of disease genes, spectrum of mutations and implications for molecular diagnosis strategy. Circulation. 2003;107:2227–2232. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

