The helicases DinG, Rep and UvrD cooperate to promote replication across transcription units in vivo
- PMID: 19851282
- PMCID: PMC2770101
- DOI: 10.1038/emboj.2009.308
The helicases DinG, Rep and UvrD cooperate to promote replication across transcription units in vivo
Erratum in
- EMBO J. 2010 Jan 6;29(1):278
Abstract
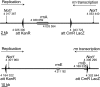
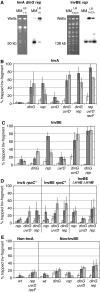
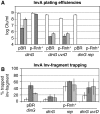
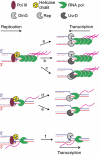
How living cells deal with head-on collisions of the replication and transcription complexes has been debated for a long time. Even in the widely studied model bacteria Escherichia coli, the enzymes that take care of such collisions are still unknown. We report here that in vivo, the DinG, Rep and UvrD helicases are essential for efficient replication across highly transcribed regions. We show that when rRNA operons (rrn) are inverted to face replication, the viability of the dinG mutant is affected and over-expression of RNase H rescues the growth defect, showing that DinG acts in vivo to remove R-loops. In addition, DinG, Rep and UvrD exert a common function, which requires the presence of two of these three helicases. After replication blockage by an inverted rrn, Rep in conjunction with DinG or UvrD removes RNA polymerase, a task that is fulfilled in its absence by the SOS-induced DinG and UvrD helicases. Finally, Rep and UvrD also act at inverted sequences other than rrn, and promote replication through highly transcribed regions in wild-type E. coli.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


References
-
- Bartlett MS, Gaal T, Ross W, Gourse RL (1998) RNA polymerase mutants that destabilize RNA polymerase-promoter complexes alter NTP-sensing by rrn P1 promoters. J Mol Biol 279: 331–345 - PubMed
-
- Bedinger P, Hochstrasser M, Jongeneel CV, Alberts BM (1983) Properties of the T4 bacteriophage DNA replication apparatus: the T4 dda DNA helicase is required to pass a bound RNA polymerase molecule. Cell 34: 115–123 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

