'Edgetic' perturbation of a C. elegans BCL2 ortholog
- PMID: 19855391
- PMCID: PMC2865203
- DOI: 10.1038/nmeth.1394
'Edgetic' perturbation of a C. elegans BCL2 ortholog
Erratum in
- Nat Methods. 2009 Dec;6(12):935
Abstract
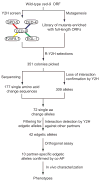
Genes and gene products do not function in isolation but within highly interconnected 'interactome' networks, modeled as graphs of nodes and edges representing macromolecules and interactions between them, respectively. We propose to investigate genotype-phenotype associations by methodical use of alleles that lack single interactions, while retaining all others, in contrast to genetic approaches designed to eliminate gene products completely. We describe an integrated strategy based on the reverse yeast two-hybrid system to isolate and characterize such edge-specific, or 'edgetic', alleles. We established a proof of concept with CED-9, a Caenorhabditis elegans BCL2 ortholog. Using ced-9 edgetic alleles, we uncovered a new potential functional link between apoptosis and a centrosomal protein. This approach is amenable to higher throughput and is particularly applicable to interactome network analysis in organisms for which transgenesis is straightforward.
Figures


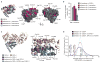


Comment in
-
You too can play with an edge.Nat Methods. 2009 Nov;6(11):797-8. doi: 10.1038/nmeth1109-797. Nat Methods. 2009. PMID: 19876015 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

