The Arabidopsis thaliana double-stranded RNA binding protein DRB1 directs guide strand selection from microRNA duplexes
- PMID: 19861421
- PMCID: PMC2779670
- DOI: 10.1261/rna.1646909
The Arabidopsis thaliana double-stranded RNA binding protein DRB1 directs guide strand selection from microRNA duplexes
Abstract
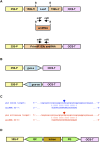
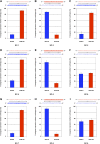
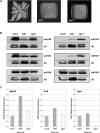
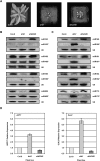
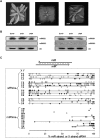
In Arabidopsis thaliana (Arabidopsis), DICER-LIKE1 (DCL1) functions together with the double-stranded RNA binding protein (dsRBP), DRB1, to process microRNAs (miRNAs) from their precursor transcripts prior to their transfer to the RNA-induced silencing complex (RISC). miRNA-loaded RISC directs RNA silencing of cognate mRNAs via ARGONAUTE1 (AGO1)-catalyzed cleavage. Short interefering RNAs (siRNAs) are processed from viral-derived or transgene-encoded molecules of double-stranded RNA (dsRNA) by the DCL/dsRBP partnership, DCL4/DRB4, and are also loaded to AGO1-catalyzed RISC for cleavage of complementary mRNAs. Here, we use an artificial miRNA (amiRNA) technology, transiently expressed in Nicotiana benthamiana, to produce a series of amiRNA duplexes with differing intermolecular thermostabilities at the 5' end of duplex strands. Analyses of amiRNA duplex strand accumulation and target transcript expression revealed that strand selection (amiRNA and amiRNA*) is directed by asymmetric thermostability of the duplex termini. The duplex strand possessing a lower 5' thermostability was preferentially retained by RISC to guide mRNA cleavage of the corresponding target transgene. In addition, analysis of endogenous miRNA duplex strand accumulation in Arabidopsis drb1 and drb2345 mutant plants revealed that DRB1 dictates strand selection, presumably by directional loading of the miRNA duplex onto RISC for passenger strand degradation. Bioinformatic and Northern blot analyses of DCL4/DRB4-dependent small RNAs (miRNAs and siRNAs) revealed that small RNAs produced by this DCL/dsRBP combination do not conform to the same terminal thermostability rules as those governing DCL1/DRB1-processed miRNAs. This suggests that small RNA processing in the DCL1/DRB1-directed miRNA and DCL4/DRB4-directed sRNA biogenesis pathways operates via different mechanisms.
Figures

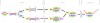
Similar articles
-
DRB2 is required for microRNA biogenesis in Arabidopsis thaliana.PLoS One. 2012;7(4):e35933. doi: 10.1371/journal.pone.0035933. Epub 2012 Apr 24. PLoS One. 2012. PMID: 22545148 Free PMC article.
-
SINE RNA induces severe developmental defects in Arabidopsis thaliana and interacts with HYL1 (DRB1), a key member of the DCL1 complex.PLoS Genet. 2008 Jun 13;4(6):e1000096. doi: 10.1371/journal.pgen.1000096. PLoS Genet. 2008. PMID: 18551175 Free PMC article.
-
DRB2, DRB3 and DRB5 function in a non-canonical microRNA pathway in Arabidopsis thaliana.Plant Signal Behav. 2012 Oct 1;7(10):1224-9. doi: 10.4161/psb.21518. Epub 2012 Aug 20. Plant Signal Behav. 2012. PMID: 22902697 Free PMC article.
-
Plant dicer-like proteins: double-stranded RNA-cleaving enzymes for small RNA biogenesis.J Plant Res. 2017 Jan;130(1):33-44. doi: 10.1007/s10265-016-0877-1. Epub 2016 Nov 24. J Plant Res. 2017. PMID: 27885504 Review.
-
microRNA strand selection: Unwinding the rules.Wiley Interdiscip Rev RNA. 2021 May;12(3):e1627. doi: 10.1002/wrna.1627. Epub 2020 Sep 20. Wiley Interdiscip Rev RNA. 2021. PMID: 32954644 Free PMC article. Review.
Cited by
-
Small RNA pathways and diversity in model legumes: lessons from genomics.Front Plant Sci. 2013 Jul 10;4:236. doi: 10.3389/fpls.2013.00236. eCollection 2013. Front Plant Sci. 2013. PMID: 23847640 Free PMC article.
-
Differential mRNA Accumulation upon Early Arabidopsis thaliana Infection with ORMV and TMV-Cg Is Associated with Distinct Endogenous Small RNAs Level.PLoS One. 2015 Aug 3;10(8):e0134719. doi: 10.1371/journal.pone.0134719. eCollection 2015. PLoS One. 2015. PMID: 26237414 Free PMC article.
-
Genome-Wide Association Study of Wood Anatomical and Morphological Traits in Populus trichocarpa.Front Plant Sci. 2020 Sep 9;11:545748. doi: 10.3389/fpls.2020.545748. eCollection 2020. Front Plant Sci. 2020. PMID: 33013968 Free PMC article.
-
The Complexity of Posttranscriptional Small RNA Regulatory Networks Revealed by In Silico Analysis of Gossypium arboreum L. Leaf, Flower and Boll Small Regulatory RNAs.PLoS One. 2015 Jun 12;10(6):e0127468. doi: 10.1371/journal.pone.0127468. eCollection 2015. PLoS One. 2015. PMID: 26070200 Free PMC article.
-
DRB2 is required for microRNA biogenesis in Arabidopsis thaliana.PLoS One. 2012;7(4):e35933. doi: 10.1371/journal.pone.0035933. Epub 2012 Apr 24. PLoS One. 2012. PMID: 22545148 Free PMC article.
References
-
- Adenot X, Elmayan T, Lauressergues D, Boutet S, Bouché N, Gasciolli V, Vaucheret H. DRB4-dependent TAS3 trans-acting siRNAs control leaf morphology through AGO7. Curr Biol. 2006;16:927–932. - PubMed
-
- Allen E, Xie Z, Gustafson AM, Carrington JC. MicroRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell. 2005;121:207–221. - PubMed
-
- Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R, et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science. 2003;301:653–657. - PubMed
-
- Bagga S, Bracht J, Hunter S, Massirer K, Holtz J, Eachus R, Pasquinelli AE. Regulation by let-7 and lin-4 miRNAs results in target mRNA degradation. Cell. 2005;122:553–563. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
