Kinetics of stop codon recognition by release factor 1
- PMID: 19874047
- PMCID: PMC2789991
- DOI: 10.1021/bi901577d
Kinetics of stop codon recognition by release factor 1
Abstract
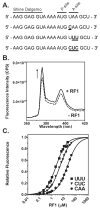
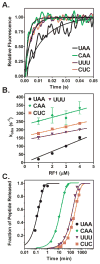
Recognition of stop codons by class I release factors is a fundamental step in the termination phase of protein synthesis. Since premature termination is costly to the cell, release factors have to efficiently discriminate between stop and sense codons. To understand the mechanism of discrimination between stop and sense codons, we developed a new, pre-steady state kinetic assay to monitor the interaction of RF1 with the ribosome. Our results show that RF1 associates with similar association rate constants with ribosomes programmed with stop or sense codons. However, dissociation of RF1 from sense codons is as much as 3 orders of magnitude faster than from stop codons. Interestingly, the affinity of RF1 for ribosomes programmed with different sense codons does not correlate with the defects in peptide release. Thus, discrimination against sense codons is achieved with both an increase in the dissociation rates and a decrease in the rate of peptide release. These results suggest that sense codons inhibit conformational changes necessary for RF1 to stably bind to the ribosome and catalyze peptide release.
Figures
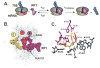
Similar articles
-
Structural aspects of translation termination on the ribosome.RNA. 2011 Aug;17(8):1409-21. doi: 10.1261/rna.2733411. Epub 2011 Jun 23. RNA. 2011. PMID: 21700725 Free PMC article. Review.
-
Thermodynamic and kinetic insights into stop codon recognition by release factor 1.PLoS One. 2014 Apr 3;9(4):e94058. doi: 10.1371/journal.pone.0094058. eCollection 2014. PLoS One. 2014. PMID: 24699820 Free PMC article.
-
Mechanism of premature translation termination on a sense codon.J Biol Chem. 2018 Aug 10;293(32):12472-12479. doi: 10.1074/jbc.AW118.003232. Epub 2018 Jun 25. J Biol Chem. 2018. PMID: 29941456 Free PMC article.
-
Structural basis for stop codon recognition in eukaryotes.Nature. 2015 Aug 27;524(7566):493-496. doi: 10.1038/nature14896. Epub 2015 Aug 5. Nature. 2015. PMID: 26245381 Free PMC article.
-
Crystal structures of 70S ribosomes bound to release factors RF1, RF2 and RF3.Curr Opin Struct Biol. 2012 Dec;22(6):733-42. doi: 10.1016/j.sbi.2012.08.004. Epub 2012 Sep 19. Curr Opin Struct Biol. 2012. PMID: 22999888 Free PMC article. Review.
Cited by
-
Single-molecule fluorescence measurements of ribosomal translocation dynamics.Mol Cell. 2011 May 6;42(3):367-77. doi: 10.1016/j.molcel.2011.03.024. Mol Cell. 2011. PMID: 21549313 Free PMC article.
-
Diversity and Similarity of Termination and Ribosome Rescue in Bacterial, Mitochondrial, and Cytoplasmic Translation.Biochemistry (Mosc). 2021 Sep;86(9):1107-1121. doi: 10.1134/S0006297921090066. Biochemistry (Mosc). 2021. PMID: 34565314 Free PMC article. Review.
-
Structural aspects of translation termination on the ribosome.RNA. 2011 Aug;17(8):1409-21. doi: 10.1261/rna.2733411. Epub 2011 Jun 23. RNA. 2011. PMID: 21700725 Free PMC article. Review.
-
Thermodynamic and kinetic insights into stop codon recognition by release factor 1.PLoS One. 2014 Apr 3;9(4):e94058. doi: 10.1371/journal.pone.0094058. eCollection 2014. PLoS One. 2014. PMID: 24699820 Free PMC article.
-
Translational termination without a stop codon.Science. 2016 Dec 16;354(6318):1437-1440. doi: 10.1126/science.aai9127. Epub 2016 Dec 1. Science. 2016. PMID: 27934701 Free PMC article.
References
-
- Brenner S, Stretton AO, Kaplan S. Genetic code: the ‘nonsense’ triplets for chain termination and their suppression. Nature. 1965;206:994–998. - PubMed
-
- Konecki DS, Aune KC, Tate W, Caskey CT. Characterization of reticulocyte release factor. J Biol Chem. 1977;252:4514–4520. - PubMed
-
- Jorgensen F, Adamski FM, Tate WP, Kurland CG. Release factor-dependent false stops are infrequent in Escherichia coli. J Mol Biol. 1993;230:41–50. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

