The photosystem II light-harvesting protein Lhcb3 affects the macrostructure of photosystem II and the rate of state transitions in Arabidopsis
- PMID: 19880802
- PMCID: PMC2782274
- DOI: 10.1105/tpc.108.064006
The photosystem II light-harvesting protein Lhcb3 affects the macrostructure of photosystem II and the rate of state transitions in Arabidopsis
Abstract
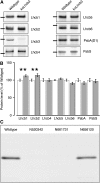
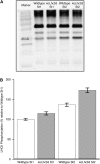
The main trimeric light-harvesting complex of higher plants (LHCII) consists of three different Lhcb proteins (Lhcb1-3). We show that Arabidopsis thaliana T-DNA knockout plants lacking Lhcb3 (koLhcb3) compensate for the lack of Lhcb3 by producing increased amounts of Lhcb1 and Lhcb2. As in wild-type plants, LHCII-photosystem II (PSII) supercomplexes were present in Lhcb3 knockout plants (koLhcb3), and preservation of the LHCII trimers (M trimers) indicates that the Lhcb3 in M trimers has been replaced by Lhcb1 and/or Lhcb2. However, the rotational position of the M LHCII trimer was altered, suggesting that the Lhcb3 subunit affects the macrostructural arrangement of the LHCII antenna. The absence of Lhcb3 did not result in any significant alteration in PSII efficiency or qE type of nonphotochemical quenching, but the rate of transition from State 1 to State 2 was increased in koLhcb3, although the final extent of state transition was unchanged. The level of phosphorylation of LHCII was increased in the koLhcb3 plants compared with wild-type plants in both State 1 and State 2. The relative increase in phosphorylation upon transition from State 1 to State 2 was also significantly higher in koLhcb3. It is suggested that the main function of Lhcb3 is to modulate the rate of state transitions.
Figures

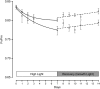
References
-
- Ahn, T.K., Avenson, T.J., Ballottari, M., Cheng, Y.-C., Niyogi, K.K., Bassi, R., and Fleming, G.R. (2008). Architecture of a charge-transfer state regulating light harvesting in a plant antenna protein. Science 320 794–797. - PubMed
-
- Allen, J.F. (2003). State transitions - A question of balance. Science 299 1530–1532. - PubMed
-
- Allen, J.F. (2005). Photosynthesis: The processing of redox signals in chloroplasts. Curr. Biol. 15 929–932. - PubMed
-
- Allen, J.F., and Forsberg, J. (2001). Molecular recognition in thylakoid structure and function. Trends Plant Sci. 6 317–326. - PubMed
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

